|
Research Ideas and Outcomes :
Research Article
|
|
Corresponding author: Steen Dupont (steen.dupont@nhm.ac.uk)
Received: 24 Apr 2020 | Published: 27 Apr 2020
© 2020 Steen Dupont, Josh Humphries, Alice Butcher, Edward Baker, Laura Balcells, Benjamin Price
This is an open access article distributed under the terms of the Creative Commons Attribution License (CC BY 4.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Dupont S, Humphries J, Butcher AJ, Baker E, Balcells L, Price BW (2020) Ahead of the curve: three approaches to mass digitisation of vials with a focus on label data capture. Research Ideas and Outcomes 6: e53606. https://doi.org/10.3897/rio.6.e53606
|
|
Abstract
There has been little research on novel approaches to digitising liquid-preserved natural history specimens stored in jars or vials. This paper discusses and analyses three different prototypes for high-throughput digitisation using cheap, readily available components. This paper has been written for other digitisation teams or curators who want to trial or improve upon these new digitisation approaches in liquid preserved collections.
Keywords
natural history collections, imaging, vials, jars, liquid preserved collections, spirit collections, wet specimens, digitisation
Introduction
A large proportion of natural history collections are stored in jars or vials, for example a recent inventory of the Natural History Museum, London (NHM) collections estimates over 17 million specimens are stored in 1.8 million curatorial units (
Previously published workflows for imaging wet specimens and their labels have removed the specimens from their container to be laid out for imaging in a petri dish (
Recent advances in multi-camera imaging, focussed on label data extraction from pinned insects (
MALICE: Mirror Angled Label Image Capture and Extraction
The MALICE setup is an iteration of the ALICE system described in
Hardware
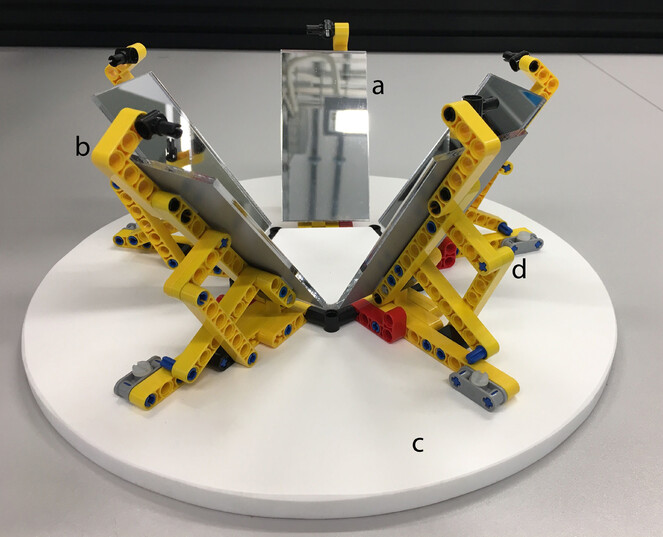
Built with materials including LEGO©, 5 acrylic mirrors, a base and specimen holder inset cut from Formex (Fig.
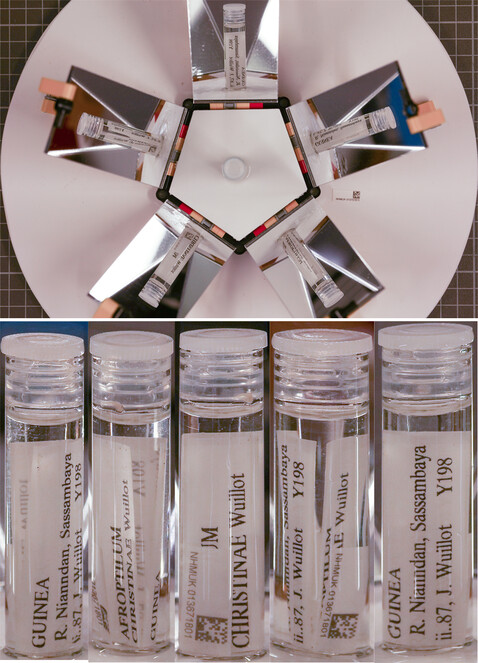
Workflow
Imaging: A vial is removed from the jar, placed in the centre of the MALICE setup, with a unique identifier (UID) label, encoded as a barcode, placed horizontally next to it. A single image is taken from above,capturing five views of the vial in the mirror images. The vial is then opened andthe barcode label is inserted. The vial is then re-sealed and placed back in the original jar.
Processing:
- Images were renamed with Gouda, using the command: "decode_barcodes.exe -a rename --avoid-collisions libdmtx "path to files"".
- The five vial reflections were cropped out of the image using Adobe Photoshop (CC 2019).
- Cropped images were manually deskewed, rotated and reflected using Photoshop creating a composite image.
VILE: Vial Imaging and Label Extraction
Successfully taking multiple images by rotating an object in front of a fixed camera is a straightforward approach, but requires some automation to make the approach scalable for mass digitisation. VILE uses an off-the-shelf rotary table and controller positioned in front of a consumer camera, controlled using a laptop.
Hardware
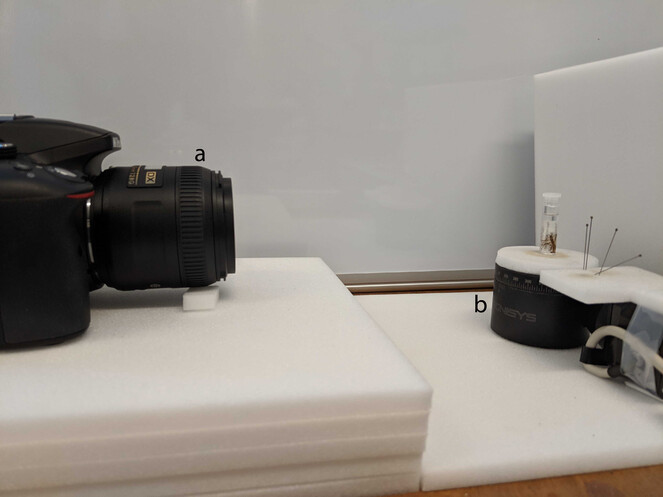
Images were taken using a Nikon D5300 camera. Specimen vials were placed on a Stackshot rotary table with controller (https://www.cognisys-inc.com/store/rotary-table.html) (Fig.
Workflow
Imaging: A vial is removed from the jar, placed in the centre of the rotating stage, with a UID label, encoded as a barcode placed vertically on a stationary stage within the imaging frame. Images are taken at six different angles corresponding to 0, 72, 144, 216, 288 and 360 degrees. The vial is then opened, the UID label inserted, the vial re-sealed and placed back in the original jar. The arbitrary degree between the images reflects a mistaken configuration of the stackshot turntable resulting in the duplication of the initial view with that of 360 degrees, which was subsequently deleted. Future imaging should utilise 0, 60, 120, 180, 240, 300 degrees to maximise label views while removing duplication.
Processing: Image processing followed these steps:
- Images were renamed with Gouda, using the command: "decode_barcodes.exe -a rename --avoid-collisions libdmtx "path to files"".
- Images were cropped, minimizing extra space, with XNConvert v1.76, saving the output images as JPG at 90% quality.
- Stitch 5 images together using mageMagick 7.0.7 ( https://github.com/ImageMagick ) with a recursive script based on the image name (e.g. “magick 013670425-4.jpg 013670425-3.jpg 013670425-2.jpg 013670425-1.jpg 013670425.jpg +append 013670425_combined.jpg”)
ReVILE: Revolving Vial Imaging and Label Extraction
The technique applied in the ReVILE setup is based on the 19th century concept of slit-scanning and primarily used today for panoramic photography. This concept involves a slit moving across the photographic medium (film or digital sensor), thereby exposing only a small section of the photographic medium to light at any given time. The implication of the temporal element of the slit scan means that a slit-scan image is composed of several snapshots in time. By moving the subject of the photograph instead of the slit, the process is inverted, reproducing a 3D object on a 2D plane. The imaging concept of ReVILE is no different from VILE in that an object is rotated in front of a stationary camera. However, generating an image by extracting vertical lines from the camera video and aligning these to create a composite image requires a more precise camera-object alignment and a rotation speed that is determined by the diameter of the object.
Hardware
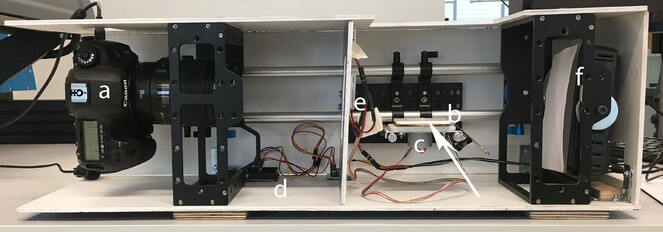
Camera, turntable, controller and lighting were built into a formex casing. The setup presented uses a Canon 5DsR camera with a 100mm Canon macro lens for capturing video and still images. The turntable is custom-built using 2mm aluminum, and polystyrene sheets. Movement of the turntable is provided by a 5v stepper motor (28BYJ-48 5V) controlled by a drive chip (ULN2003) and an Arduino UNO. Lighting was provided by 3 COB 48-SMD LED panels for front lighting and a Neewer 40004082 light panel for backlighting (Fig.
Workflow
Imaging: A vial is placed in the centre of the turntable and filmed over a 720 degree rotation. Rotating the vial multiple times ensures that all possible angles are covered. The setup is calibrated by taking a single frame of a scale at the centre of the turntable and using image software to estimate the width (in pixels) of a single millimeter. The number of frames \(n_f\) required can then be calculated as such:
\(n_f = {2\pi dlxw_1 \over 2lx-dsw_1}\)
Where \(d\) is the vial diameter (in mm), \(l\) is the focal length (in mm), \(x\) is the size of the frame (in pixels) across the x axis of the vial (i.e. width if landscape, height if portrait), \(s\) is the size of the sensor (in mm) across that same x dimension, and \(w_1\) is the width of 1mm in pixels (obtained during the calibration step).
The duration of a single rotation can be calculated by dividing this figure by the stream's framerate: \(t = {n_f \over r}\)
This figure is largely a helpful estimate and high levels of accuracy are not necessary; it is preferable to err on the side of longer rotations to ensure that detail is not missed.
In the example dataset, videos were captured at approximately 60 frames per second, at a focal length of 100mm and a resolution of 1280x720 pixels (in portrait orientation). The size of the sensor on the Canon 5DSR is 36x24mm. The width of 1mm on the calibration frame was approximately 21 pixels.
For a 10mm vial: \(n_f = {2\pi \times 10 \times 100 \times 720 \times 21 \over 2 \times 100 \times 720 - 10 \times 24 \times 21 } = 684\) and \(t = {684 \over 60} = 11.4\)
For a 20mm vial: \(n_f = {2\pi \times 20 \times 100 \times 720 \times 21 \over 2 \times 100 \times 720 - 20 \times 24 \times 21 } = 1419\) and \(t = { 1419 \over 60} = 23.7\)
Processing: ReVILE is capable of producing 3 separate image outputs outlined in the results section. The primary output is the composite rolled out image. The workflow of the rollout photography output is outlined below and the potential for the other two outputs - a composite image of the vial at different angles (equivalent to the VILE output) and an interactive digital-only output that allows for user rotation of the imaged vial - are examined in the discussion.
Custom software was developed to control the camera and turntable as well as process the camera’s outputs to produce the rolled out image of the object. This software is available at https://github.com/NaturalHistoryMuseum/revile.
Once the object has been securely positioned on the turntable in front of the camera, the software follows the following steps to produce the rolled out image of the object:
- Initialise the video capture process - this is done first as it takes a couple of seconds to begin capturing, but step 2 should generally begin first;
- Start rotating the turntable for the specified number of seconds;
- Iterate through the stream or video frames, extracting the middle column or row (depending on camera orientation) of pixels from each frame;
- Concatenate the extracted lines of pixels from each frame together to build the image;
- Undertake any postprocessing steps, e.g. rotating the output;
- Crop the image so that it shows only one complete rotation, using feature matching to locate similar columns of pixels; and
- Write two JPEG images to disk: the complete, "raw" output, and the cropped version.
Using the hardware setup pictured in Fig.
The speed of the turntable’s rotation is determined by the diameter of the object being processed. Wider objects, such as a jar, require a slower rotation while thinner objects, such as a vial, can be rotated faster. The software provides an estimate function through its command line interface (CLI) which can be used to guess the rotation duration for the object given its diameter in mm, number of pixels per mm as seen by the camera (by default, 21) and the source frame rate (either the stream frame rate or the video frame rate).
Data resources
Example media from the MALICE, VILE and ReVILE setups can be downloaded here:
MALICE:https://doi.org/10.5281/zenodo.3497060
VILE: https://doi.org/10.5281/zenodo.3462015
ReVILE: https://doi.org/10.5281/zenodo.3727644
The software for ReVILE can be found here:
Results
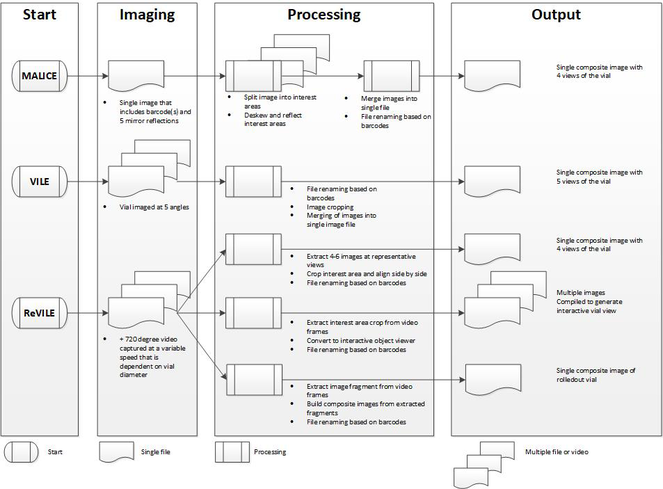
The workflow for each of the three approaches is outlined in Fig.
Results of the imaging tests completed for the three setups including test metrics and an estimate of the imaging time required for a collection of 5000 vials for reference. Note the estimates for MALICE and ReVILE exclude any recuration and rehousing.
| Number of vials imaged | Total imaging time (minutes) | Еstimated vials/ person hour | Estimated imaging time in person days (5.5 hrs) for a vial collection of n = 5000 with databasing | |
|---|---|---|---|---|
| MALICE |
42 (only imaged) |
15 |
168 |
23.9 |
| 48 (with databasing) | 75 | 38 | ||
| VILE | 2349 (with databasing) | Not recorded | 30-40 | 26.5 |
| ReVILE |
21 (only imaged) |
18 |
70 |
31* |
| (with databasing) | 29* |
*assumed increase in imaging time of 72 seconds per vile when databasing based on timings from MALICE, time also includes image processing.
ReVILE: Two Rollout photography outputs of ReVILE. Left to right: a frame from the video (rotated 90° clockwise) showing the vial itself; the uncropped rollout image, covering more than one full rotation; the cropped rollout image, showing only one 360° rotation; the cropped rollout image, "shifted" across (by transferring a manually-defined block of pixel columns from the left side of the image to the right) to show the complete label
Conclusions
The three setups described provide means of digitising wet collections at a considerable speed and with a variation of outputs that are useful. We propose that a ReVILE approach is a more versatile solution for wet collection digitisation. While MALICE is arguably the fastest approach the setup is not ameanable to the large variation of vials available in collections not only due to mirror size requirements, but also the limits to the depth of field required by the camera. Using a combination of the VILE and ReVILE approach which have the same setup but differ in the image capture and processing it is possible to create the output of both setups as is suggested in Fig.
A summary of the pros and cons of the three systems presented.
| Pros | Cons | |
|---|---|---|
| MALICE |
|
|
| VILE |
|
|
| ReVILE |
|
|
Acknowledgements
We would like to thank Laura Sivess and Lyndall Pereira da Conceicoa for discussion around the VILE setup. Zoe Hughes and Jennifer White for testing the ReVILE setup and Vladimir Blagoderov for valuable discussion and suggestions for the MALICE design. In addition we would like to thank Laurence Livermore for constructive comments on the manuscript.
Funding program
Funding for staff time and materials to create ReVILE and MALICE was provided by the Natural History Museum Digital Collections Programme and ICEDIG.
ICEDIG was funded under H2020-EU.1.4.1.1. - Developing new world-class research infrastructures.
Grant title
ICEDIG – “Innovation and consolidation for large scale digitisation of natural heritage”, Grant Agreement No. 777483.
Author contributions
All authors contributed to devising the project. Software was developed by Alice Butcher with additional contributions by Josh Humphries. All authors contributed to writing the manuscript.
References
-
Wet collections accession: a workflow based on a large stonefly (Insecta, Plecoptera) donation.Biodiversity Data Journal6:e30256. https://doi.org/10.3897/bdj.6.e30256
-
Imaging natural history museum collections from the bottom up: 3D print technology facilitates imaging of fluid-stored arthropods with flatbed scanners.ZooKeys795:49‑65. https://doi.org/10.3897/zookeys.795.28416
-
ALICE: Angled Label Image Capture and Extraction for high throughput insect specimen digitisation.OSF Preprintshttps://doi.org/10.31219/osf.io/s2p73
-
The Natural History Museum Data Portal.Database: the journal of biological databases and curation2019https://doi.org/10.1093/database/baz038
-
Join the Dots: assessing 80 million items at the Natural History Museum, London.Biodiversity Information Science and Standards2:е26500. https://doi.org/10.3897/biss.2.26500