|
Research Ideas and Outcomes :
Short Communication
|
|
Corresponding author: Felicitas Löffler (felicitas.loeffler@uni-jena.de)
Received: 26 Apr 2021 | Published: 26 May 2021
© 2021 Felicitas Löffler, Andreas Schuldt, Birgitta König-Ries, Helge Bruelheide, Friederike Klan
This is an open access article distributed under the terms of the Creative Commons Attribution License (CC BY 4.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation:
Löffler F, Schuldt A, König-Ries B, Bruelheide H, Klan F (2021) A Test Collection for Dataset Retrieval in Biodiversity Research. Research Ideas and Outcomes 7: e67887. https://doi.org/10.3897/rio.7.e67887
|
|
Abstract
Searching for scientific datasets is a prominent task in scholars' daily research practice. A variety of data publishers, archives and data portals offer search applications that allow the discovery of datasets. The evaluation of such dataset retrieval systems requires proper test collections, including questions that reflect real world information needs of scholars, a set of datasets and human judgements assessing the relevance of the datasets to the questions in the benchmark corpus. Unfortunately, only very few test collections exist for a dataset search. In this paper, we introduce the BEF-China test collection, the very first test collection for dataset retrieval in biodiversity research, a research field with an increasing demand in data discovery services. The test collection consists of 14 questions, a corpus of 372 datasets from the BEF-China project and binary relevance judgements provided by a biodiversity expert.
Keywords
dataset search, dataset retrieval, test collection, biodiversity research
Introduction
Dataset search and data reuse are becoming more important in scholars' research practice. Instead of recreating datasets by repeating experiments or for the comparison of new datasets with similar data collected under different conditions, scholars increasingly search for existing datasets. For example, GBIF's scientific report (
One research domain with an increasing demand for data discovery services is biodiversity research, a domain that examines the variety of species, their genetic diversity and ecological diversity. Scholars working in the fields of biodiversity research often need to search and combine several datasets from different experiments to answer a research question. Hence, proper data retrieval systems are needed to support these data discovery tasks. In this work, we introduce the first test collection for dataset retrieval in biodiversity research. We focus on an important sub-domain in biodiversity research, ecosystem functioning, that has been intensively studied in the BEF-China project (https://www.bef-china.com). In this project, 372 datasets are publicly available with structured metadata files. Metadata are descriptive information about the measured or observed primary data and contain information such as author, collection time, title, abstract, keywords and parameters measured. Depending on the domain, metadata are provided in a specific structure or metadata schema. In the BEF-China project, all metadata files are provided in EML, the Ecological Metadata Language (KNB (ecoinformatics.org)). Providing relevance judgements is a very time-consuming task. Therefore, we only selected 14 questions collected in various biodiversity projects. They do not cover all search interests in biodiversity research, but reflect real world information needs of scholars. Binary human relevance assessments are provided by a biodiversity expert.
The structure of the paper is as follows: at first, we present related work. Afterwards, we describe the creation steps of the BEF-China test collection, including data collection, question collection and human ratings. At the end, we conclude with a summary of our findings.
The test collection is publicly available on GitHub at https://github.com/fusion-jena/befchina-test-collection and on Zenodo via http://doi.org/10.5281/zenodo.4704947.
Related Work
A retrieval system consists of a collection of documents (a corpus) and a user’s information needs that are described by a set of keywords (query). The main aim of the retrieval process is to return a ranked list of documents that match the user’s query. Numerous evaluation measures have been developed to assess the effectiveness of retrieval systems in terms of relevance. For this purpose, a test collection is required that consists of three parts (
- a corpus of documents,
- representative information needs expressed as queries and
- a set of relevance judgements provided by human judges containing assessments of the relevance of a document for given queries.
If judgements are available for the entire corpus, they serve as baseline (“gold standard”) and can be used to determine the fraction of relevant documents a search system finds for a specific query.
In Life Sciences, one of the first Information Retrieval benchmarks is the Genomics Track Challenge (
The BioCADDIE Test Collection (
To the best of our knowledge, there is no test collection available for dataset search in biodiversity research. Therefore, in the following, we introduce our test collection for dataset retrieval in biodiversity research.
The BEF-China Dataset Retrieval Test Collection
Biodiversity research nowadays is a very heterogenous research field that goes beyond the exploration of species richness and taxon relations. Over the last few decades, research into the relationships between biodiversity and ecosystem functioning and the consequences of biodiversity change for ecosystems, has become a key topic of interdisciplinary biodiversity research (
Data Collection
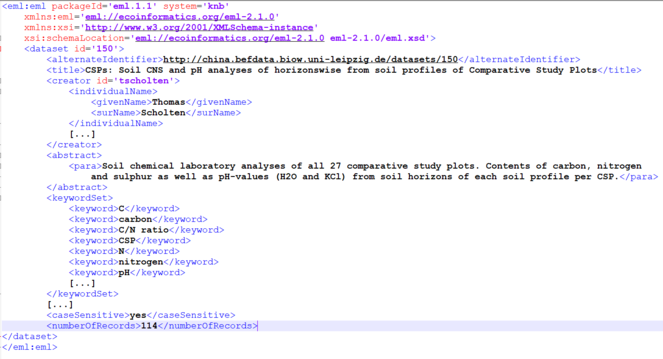
The data collected in the BEF-China project are publicly provided in a corpus of 372 metadata files. Most datasets also provide open access to the primary data. The metadata information are stored in XML files following the EML metadata schema (https://eml.ecoinformatics.org/). A data manager supported the scientists in providing proper data descriptions to ensure FAIR data and metadata (
Question Collection
The development of the test collection is driven by two requirements: we aim at providing a test collection reflecting real world information needs from biodiversity scholars. At the same time, we need to ensure that at least a fraction of the datasets in the corpus is relevant to the information needs expressed in the queries. Therefore, we selected six questions from a question corpus, collected in our previous research (
BEF-China question corpus and number of datasets being relevant to a question.
| Question Number | Question | Number of datasets being relevant to this question |
| Q1 | Name 3 species that occur in the shrub layer. | 16 |
| Q2 | Find 3 plant species where root lengths (depth) have been considered | 1 |
| Q3 | Find 3 datasets from oaks where nitrogen content have been measured. | 18 |
| Q4 | Find 3 datasets where dry weights from conifers have been measured. | 5 |
| Q5 | Which nutrients occur in soil? | 20 |
| Q6 | Identify all parameters that are correlated to soil depth. | 24 |
| Q7 | Which taxa associated with tree species have been found, for example, insects on host trees? | 46 |
| Q8 | Which soil samples in BEF-China data show a low pH value? | 6 |
| Q9 | Does tree diversity reduce competition? | 40 |
| Q10 | Do the soil carbon concentrations increase with soil depth? | 6 |
| Q11 | Are there data about the leaf area index (LAI) and, in particular, in combination with diversity? | 8 |
| Q12 | How has tree height been measured in BEF-China experiments? | 25 |
| Q13 | How does the nitrogen cycle interact with water? | 20 |
| Q14 | How significant is the role of throughfall as water input to the forest floor? | 4 |
Human Assessments
Human assessments are required to determine which dataset is relevant to which question. This assessment was provided by one of the co-authors who was the data manager of the BEF-China project at this time and who has acquired a comprehensive overview of the entire corpus of datasets. For each question, he evaluated whether a dataset is relevant or not relevant to each of the 14 questions. He was asked to judge a dataset also as 'relevant' if it only partially comprised relevant data. As the corpus also contains presentations, plot descriptions and theses established in the scope of the BEF-China project, not all datasets are relevant to one of the 14 questions. However, the biodiversity expert took the necessary time to go through all 372 datasets per question. Hence, 5208 relevance judgements (14 questions x 372 datasets) had to be conducted. Out of these 5208 relevance judgements, 239 were judged as relevant or partially relevant. These relevance judgements are provided in a txt file complying with the TREC benchmark data format. An entry in the txt file looks as follows:
1::161::1::1424380312
The first number denotes the question number, the second number provides the dataset number, the third denotes the relevance judgement (1-relevant) and the last number is the timestamp of the creation of the entry. All datasets of the BEF-China corpus that are not mentioned as relevant for a question are deemed not to be relevant. Hence, the txt file only contains the relevant datasets per question.
Conclusion
In this work, we presented the first test collection for a dataset search in biodiversity research. The test collection is publicly available. In our future work, we would like to use the presented test collection for evaluating dataset retrieval systems in the biodiversity domain, such as presented in
Acknowledgements
This work was conducted in the scope of the GFBio project (DFG project number: 229241684) funded by the Deutsche Forschungsgemeinschaft (DFG). The BEF-China project (DFG FOR 891/1-3, DFG project number: 35758305) was also supported by the Deutsche Forschungsgemeinschaft (DFG). We further appreciate the support received by the Sino‐German Centre for Research Promotion (GZ 986), the German Centre for Integrative Biodiversity Research (iDiv) Halle-Jena-Leipzig (DFG - FZT 118, 202548816) and the National Natural Science Foundation of China (31870409).
References
- Community assembly during secondary forest succession in a Chinese subtropical forest.Ecological Monographs81(1):25‑41. https://doi.org/10.1890/09-2172.1
- Designing forest biodiversity experiments: general considerations illustrated by a new large experiment in subtropical China.Methods in Ecology and Evolution5(1):74‑89. https://doi.org/10.1111/2041-210X.12126
- A publicly available benchmark for biomedical dataset retrieval: the reference standard for the 2016 bioCADDIE dataset retrieval challenge.Database2017https://doi.org/10.1093/database/bax061
- GBIF Science Review 2020. https://doi.org/10.35035/bezp-jj23
- TREC genomics special issue overview.Information Retrieval12(1):1‑15. https://doi.org/10.1007/s10791-008-9076-6
- A survey of current practices in data search services.Research Data Alliance Data (RDA) Discovery Paradigms Interest Group.https://doi.org/10.17632/7j43z6n22z.1
- Honey Bee Versus Apis Mellifera: A Semantic Search for Biological Data. In:The Semantic Web: ESWC 2017 Satellite Events. ESWC 2017. Lecture Notes in Computer Science,10577.Springer, Cham,98-103pp. https://doi.org/10.1007/978-3-319-70407-4_19
- Dataset Search In Biodiversity Research: Do Metadata In Data Repositories Reflect Scholarly Information Needs?PlosONE. https://doi.org/10.1371/journal.pone.0246099
- Introduction to Information Retrieval.Cambridge University Press: New York[ISBN0521865719, 9780521865715] https://doi.org/10.1017/CBO9780511809071
- CSPs: Soil CNS and pH analyses of horizonswise from soil profiles of Comparative Study Plots. URL: https://data.botanik.uni-halle.de/bef-china/datasets/150
- Biodiversity and Ecosystem Functioning.Annual Review of Ecology, Evolution, and Systematics45(1):471‑493. https://doi.org/10.1146/annurev-ecolsys-120213-091917
- An overview of the BIOASQ large-scale biomedical semantic indexing and question answering competition.BMC Bioinformatics16(138). https://doi.org/10.1186/s12859-015-0564-6
- An Introduction to Question Answering over Linked Data. In:Reasoning Web. Reasoning on the Web in the Big Data Era: 10th International Summer School 2014, Athens, Greece, September 8-13, 2014. Proceedings.Springer International Publishing
- The FAIR Guiding Principles for scientific data management and stewardship.Scientific Data3(160018). https://doi.org/10.1038/sdata.2016.18