|
Research Ideas and Outcomes :
Guidelines
|
|
Corresponding author: Kristin A Briney (briney@caltech.edu)
Received: 14 Jul 2020 | Published: 27 Jul 2020
© 2020 Kristin Briney, Heather Coates, Abigail Goben
This is an open access article distributed under the terms of the Creative Commons Attribution License (CC BY 4.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Briney KA, Coates H, Goben A (2020) Foundational Practices of Research Data Management. Research Ideas and Outcomes 6: e56508. https://doi.org/10.3897/rio.6.e56508
|
|
Abstract
The importance of research data has grown as researchers across disciplines seek to ensure reproducibility, facilitate data reuse, and acknowledge data as a valuable scholarly commodity. Researchers are under increasing pressure to share their data for validation and reuse. Adopting good data management practices allows researchers to efficiently locate their data, understand it, and use it throughout all of the stages of a project and in the future. Additionally, good data management can streamline data analysis, visualization, and reporting, thus making publication less stressful and time-consuming. By implementing foundational practices of data management, researchers set themselves up for success by formalizing processes and reducing common errors in data handling, which can free up more time for research. This paper provides an introduction to best practices for managing all types of data.
Keywords
data management, research data management, data management plan, open data
Introduction
Researchers regularly lose data. Whether it is on the small scale like one scientist’s thesis data disappearing with a lost laptop or on the large scale like historically significant scientific data from NASA that was likely overwritten during a magnetic tape shortage in the 1970’s (
Well managed data is a benefit to any researcher as it requires less digging to find, less effort to understand, and less processing to prepare for collaboration, reuse, and sharing. Good data management prevents a failed hard drive or the loss of a key collaborator from ending a project or requiring re-collection of data. The primary motivation for data management, however, is that it makes research go more smoothly, allowing the researcher to focus on the problems of science rather than data adminstravia. Beyond the individual researcher, good data management can contribute to scientific communities as a whole -- improving the speed of discovery, enhancing the veracity of new findings, allowing for increased collaboration, and providing new opportunities for educational use of data. The practices described in this article are foundational and broadly applicable, though projects may also benefit from advanced data management practices and discipline-specific standards, not addressed here, that are tailored to the needs of individual projects.
Data management, as referred to in this article, is a set of processes and project management strategies that actively occur throughout a project. This is distinct from building a living data management plan (DMP), as recommended by
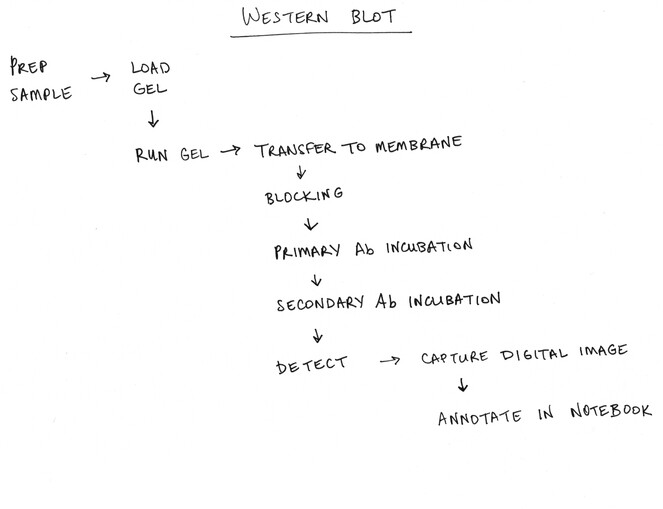
Developing a visual representation of the research process — sometimes called a workflow diagram — can be both a valuable process and tool for the integration of data management practices into research and identifying key roles and responsibilities. The process of creating this representation facilitates team learning and fosters a shared understanding of how the project, as well as functioning as a reference point for future discussions. The diagram can be as informal as a hand-drawn diagram (Fig.
This article provides ten foundational practices to improve the management of research data and files across all disciplines. While not exhaustive, this is a guide to adding small routine practices to research workflows that provide maximum impact. These practices can be applied within the standards, cultural norms, and funder requirements of individual disciplines. If you would like additional recommendations or assistance, seek out your local librarian, as academic libraries now regularly support research data management and can connect you with disciplinary resources. Much like preventative medicine, a little time spent managing data at the beginning of a research project can save a huge amount of time later in having to deal with a data disaster.
Practice 1: Keep sufficient documentation
When planning for documentation, it is useful to consider what you, your supervisor, or team members may need to know in a year when preparing for a presentation, drafting a report, or responding to questions during peer review. The most effective method for improving data management is the simplest: record the most important information for the anticipated needs. Who ran the experiment? Which procedure were used? What materials were used when they did the experiment? What were the conditions under which the observation was measured? What was the citation for the work referenced? Documentation often means the difference in being able to use an older data file or not, especially as details are forgotten over time. As funders and research communities increasingly emphasize the value of reproducibility/replication, also consider documenting additional information needed to validate or defend the results after publication.
Documentation is contained in many formats depending on the research workflow. Options include paper and electronic notebooks, README files, codebooks, data dictionaries, simple templates, etc. README files are flexible text files that provide added context, and are especially helpful in bridging the gap between physical notes and digital data (
Practice 2: Organize files and name them consistently
Organizing files is a ubiquitous challenge (
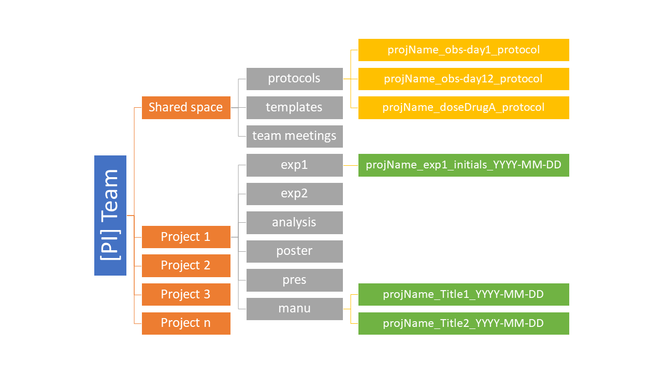
There is no ideal system for organization, just a best system for a given researcher and her data. To come up with a system, determine if there are natural groupings in the data, such as by project, analysis type, or date. Organize folders and subfolders in the most logical way that you would like to search for content; this often helps prioritize some groupings as higher level than others. Even the most comprehensive organizational structure may not fit all of the data, so a good rule of thumb is 80% covered. Note too that different types of files can have different organizational schemes. For example, you may want to organize raw data generated daily by date of collection, while data collected periodically may be naturally organized by instrument or location. However the system is created, consistent application is key. Get in the habit of always putting the data where it belongs for easy access later (Fig.
Once data is structurally organized, file naming will take searching and sorting to the next level and make it easy to distinguish between related files. A good file naming convention works for a group of related files, displays about 3 pieces of key information about any file, and is easy to visually scan (Table
|
File Type |
Sample File Name Template |
File Name Example |
|
Outputs generated from a basic science experiment |
ExperimentNumber_OutputType_Version |
Experiment25_Assay_v05.csv Experiment18_SPSSOutput_v02.tsv |
|
Manuscript drafts |
Project_Manuscript_vXX.docx |
CityHIVInc_Manuscript_v23.docx |
|
Meeting notes |
YYYYMMDD_TeamName_MeetingNotes.docx |
20181022_DDTeam_MeetingNotes.docx |
|
Literature |
Author_YYYY_ShortTitle.pdf |
Doubleday_2018_CopyrightNarrativeLitRev.pdf |
|
IRB Submission documents (sorted by order needed for IRB) |
SortingNumber_IRBType_DocumentSubject |
02_IRBExemption_MyDataSurvey |
One useful strategy for file naming and managing version controls leverages the international date convention, ISO 8601, with the format YYYY-MM-DD (or YYYYMMDD) (
Practice 3: Version the Files
Versioning, or keeping distinct copies of a document as it changes over time, is traditionally considered in the context of computer code or software, but can just as easily be applied to data, draft manuscripts, or any other file type iteratively generated during research. Procedures (either laboratory or analysis) are especially good candidates to version, as they often change over time. Applying version control to research files offers a low-barrier way to document provenance. Versioning allows a researcher to return to an earlier copy of a document, such as to rescue a portion of deleted text or to fix an analysis procedure, without having to recreate the entire thing from scratch. A Nature survey from 2015 found that 70% of researchers had tried and failed to reproduce someone else’s data and this may have been impacted by the unavailability of the procedures and document of the process (
Versioning does not need to be complex. At its most basic, versioning files means keeping an untouched copy of the original file or raw data that won’t be overwritten, separate copies of content that undergo cleaning, analytical, and visualization processes , and a clean version of the finalized file. A step above this is to periodically save new versions of a file, such as by appending a new version number like “_v03” at the end of a file name. This method is very helpful when sending paper drafts between multiple authors and allows a researcher to rescue content or analyses that appeared in an earlier version of the document. Versioning can also be done with a date (see ISO 8601 format described in Practice 2) instead of version number, depending on researcher preference. While you may consider using the marker “_FINAL” at the end of the file name only once a file will no longer receive any changes, this should be used with caution as many researchers find their work still underway or receiving revisions after peer review, leading to examples like “File_FINAL_FINAL_v6.2”. Denoting that a file is the submitted version or revised version can prevent confusing names.
The most complex, but also space saving and efficient, form of versioning is to use a version control system such as Git (https://git-scm.com), which is software designed originally for tracking code changes. While this system allows for easy sharing and tracking different branches of data as they are modified, it also comes with a steep learning curve and may not be necessary for a single project or a lone researcher.
When making choices about versioning, it is important to understand what will meet the workflow and educational demands of the research team. However, even where a simple approach is enough, always keep a master copy of the raw data in the event that processing and analysis must be redone from scratch.
Practice 4: Create a security plan, when applicable
Sensitive or confidential data — such as data about human subjects, protected intellectual property, or data regulated by privacy law or policy — will often be required to have a documented plan for how to handle that data. This makes procedure clear for anyone who interacts with that data and helps ensure that security protocols are being followed. A data security plan will be specific to the data and technology, but should include information such as: security controls, who has access to the data during the project, what happens when someone leaves the institution or needs to have their access cut off, retention period for the data, obligations to share data after the completion of the project, and how the data will be discarded or destroyed, and any requirements for training. In addition, researchers should consider the ethical responsibilities surrounding data capture and use (
At academic institutions, there are likely to be offices and resources available to assist with security. Universities will have a HIPAA compliance officer and data security officers within the information technology department and possibly individual colleges or departments. Many universities also have an information security officer or office. Beyond observing HIPAA and FERPA regulations, there may be preferred university-provided tools for encryption, classification and storage of sensitive data, and protocols. Knowing your responsibilities and using institutional resources can prevent a lack of clarity about obligation and security protocols, which has in the past resulted in a researcher being held solely accountable for an IT breach of health data (
As security needs will continue to evolve with a project, review the security plan at least annually, coordinate with institutional security officers, and update as needed.
Practice 5: Define roles and responsibilities
Much research is conducted by teams that are distributed across labs, institutions, and even nations. The roles and responsibilities of team members can be poorly understood if particular effort is not taken to agree upon and document them. Discussion and modeling best practices for research conduct are effective techniques for transmitting the values and norms of the discipline from mentor to mentee (
The onboarding process should provide an introduction to the processes and standards adopted by the team and be provided to all new members for consistency. For most, this includes team-wide elements such as compliance training, safety protocols, documentation standards (including lab notebooks), data management practices, etc. Onboarding should also include training specific to the assigned projects and a discussion of how contribution to the project is recognized, whether through authorship, acknowledgement, or some other means. The completion of this process should, of course, be documented and signed by the team lead and new team member (see Practice 1). Similarly, the exit process should provide consistent closure for a team member’s involvement with projects. This process should ensure that both the departing team member and the team retain access to the information necessary as indicated by institutional policy or funder agreement. In many cases, a trainee may continue to collaborate on manuscripts or other products after leaving the team. Continued access to the necessary data and information should be discussed and documented to enable this progression; on the other hand, immediately revoking access to all project related accounts and resources may be necessary. Updating documented roles and responsibilities quarterly, or as team members join or exit the team, is necessary for this record to be effective.
Implementing these practices within an existing team or ongoing project can be challenging. One option is to choose a new project wherein a team can implement a prenuptial roles and responsibilities agreement (
Practice 6: Back up the data
Having at least one backup can prevent myriad data headaches. Nearly everyone has personally experienced or met someone who has experienced data loss due to a hard drive crash, water damage, lost notebooks or SD cards, etc. The best way to prevent data loss is to follow the 3-2-1 Rule: three copies of the data, two geographically separate locations, and more than one type of storage device. Having multiple copies reduces the risk of corruption or loss of the master copy. Similarly, having backups in at least one other location prevents total data loss due to localized events like a lab fire or device theft. Finally, spreading data across multiple types of storage diffuses the risk inherent in different storage types; for example, between 1-2% of hard drives fail yearly with increasing failure rates for older devices (
The best backups are automated and run at a regular frequency: daily, weekly, etc. The ideal backup system and schedule are the ones that fit best into your research workflow and account for the speed and scale of data production . Once a backup system is implemented, take some time to learn how to restore data. It’s much less stressful to figure this out at the beginning than when you’re panicking over a crashed hard drive. Finally, it’s recommended to periodically confirm that backups are functioning properly to avoid unpleasant surprises when trying to recover data in a crisis.
Practice 7: Identify tool constraints
Creating a research workflow is often shaped by the available tools or the technical requirements that chosen tools have for systems, data, and interoperability. For example, many social science researchers have a preferred analytical package. Decisions about processes and tools used for data collection, storage, visualization, archiving, and sharing are primarily driven by the affordances of the analytical tool. For bench scientists, the key components of their workflow may exist outside the digital realm, while the resulting data are routinely stored in spreadsheets or databases. Whether the common component of a research workflow is a data collection instrument, a data storage platform, or an analytical package, it is helpful to consider the implications of those decisions on other aspects of the workflow.
When the primary consideration is a data collection tool, the team should consider the options for exporting data from that system or instrument, whether the data storage platform can integrate or connect directly, and whether the tools used for processing data are compatible. For example, when collecting publication metadata in XML, transforming the XML into a structured table may require use of a specific Python package. When data storage is the fixed component, considerations may include availability on mobile devices, integration with data collection and analysis tools, file size limitations, connection speeds, and file transfer rate limits. Analytical tools frequently require data to be in a specific set of structures and formats. Many specialized tools are also limited in the ways in which data can be exported and still maintain the integrity of the data (e.g., databases, NVivo or Atlas.ti, Qualtrics).
The type of data (e.g., personal health information, personally identifiable information, or other protected data) involved can also shape the workflow. For research involving data that can be made publicly available, it is still important to ensure that the system is adequately secure. This will require evaluating in the security plan (Practice 4) both individual software tools and any potential security gaps between tools. While many information security decisions are made based on risks related to legal obligations, comprehensive integrity of research data is also a concern. In the case of highly regulated and protected data, there are often a limited set of tools with adequate security controls that are approved. In these cases, it is important to plan far ahead if a project will require new software, infrastructure, or integrations between existing systems; otherwise, research can be delayed while these issues are resolved. At many institutions, research IT support and information security offices are available to help researchers think through these decisions and build an appropriately secure and feasible research workflow.
Practice 8: Close out the project
Closing out a project involves identifying, preparing, and collocating key research files so as to identify provenance, improve future retrieval, and reduce effort when preparing for handoff or data sharing. A formalized and documented end-of-project close out can occur at the conclusion of a grant, after publication of an article, after the completion of a portion of the project, or when a colleague is transitioning off the team (e.g. when students graduate). No matter the timing, the close out process enables you to revisit projects and find the most important documents as easily and quickly as possible.
There are two approaches to picking files while closing out a project. Creating master copies of all key files is typically done at the end of a project, while snapshotting is used for capturing files at defined phases of the project (e.g. between data collection and data processing). Both approaches will require the appraisal of which data to keep. This varies greatly by size and type of project as well as funder and disciplinary expectations. Refer to disciplinary standards or other identified obligations to your institution, funder, and journal in order to appropriately assess what data will need to be preserved for the long term.
Master copies are records of the project’s most important files and are typically fully processed and have undergone quality control procedures. Master copies are a part of an audit trail, which is required in clinical trials and other regulated research. Master copies may include protocols, final raw data, processed data, analytical scripts and logs, tables and figures, and copies of submitted and accepted manuscripts, posters, and presentations.
Creating a snapshot is useful for projects that are large, complex, or still in process; for instance, snapshots may be useful when developing a model or algorithm based on a dynamic dataset. Snapshots may include:
- Data collection instruments, protocols, and other key study documents
- Processed data that is ready for analysis
- Analytical scripts or procedures and resulting figures, tables, and other visualizations to be included in research outputs
- Outputs such as grant reports, posters, presentations, and articles
In addition to setting aside copies of key files, project close out is a good time to review file formats and storage media. This is because, at some point, file types and storage hardware become obsolete and make that data unreadable (
File formats exist on a spectrum of better-to-worse with respect to long-term preservation. The best formats (see the recommended format list from the
Migration, both for hardware and file formats, is another step in making your data future-friendly. Storage hardware should be updated before the hardware format falls out of regular usage and becomes difficult to read (e.g., floppy disks). File formats should be checked for upgrades (e.g., .DOC to .DOCX) and for alternatives closer to the “better” end of the format spectrum. During file format conversion, it’s a good idea to retain content in the original file format in the event that some information is lost. Migration activities can be reviewed every year or two, but are critical to ensuring that older data remains usable.
Master copies, snapshotting, using good file formats, and migration can pay off in time savings and lower stress when writing or revising manuscripts, presentations, or posters, or if someone questions the results. Following both file naming and organization conventions (Practice 2) and versioning files (Practice 3) during a project will facilitate connecting all copies of the files, should you need to dig further. Additionally, having clear roles and responsibilities (Practice 5) will indicate the person(s) responsible for maintaining the authoritative version of the master files.
Practice 9: Put the data in a repository
Beyond closing out the project with master files, a useful strategy for maintaining finalized data is to outsource its care to specialists by depositing it in a repository. Data repositories are the preferred venues for compliance with open data requirements from funders (
Depositing data into a repository is made easier by good data management practices and consists of activities such as data appraisal, selection, cleaning, documentation, and submission. The data selected for deposit will vary based on the project and field, however enough data should be provided to allow for reproduction of analysis or replication of the data capture processes. The data and documentation selected for project close out can serve as a starting point for selection for data sharing. The next step is to ensure that the selected data is clean and error free, a part of which includes documenting the data to the level that an outsider (or your future self) can pick up the data and start using it without needing extra information. More complex data may also need to be packaged with one of several tool options (
Practice 10: Write these conventions down [in a data management plan]
This would not be an article about data management without a recommendation to have a data management plan. But rather than being a required document for a funding application (
Everything is better with friends and this is especially true for data management. Research is often a collaborative effort and having all collaborators use the same consistent data conventions can save everyone time. This means a few things: conventions should be decided on as a group, documented in a common location, used consistently, and reviewed periodically. The most important conventions to share are organization and file naming, location of backups, nuances of documentation systems, and the security plan.
Under this philosophy, the data management plan is a living, working document meant to be frequently referred to and updated as necessary. Setting a schedule to review the plan will also help refresh all project members of data expectations. Decisions made under Practice 5 will suggest who should enforce DMP conventions and shepherd the updating process. Framing the DMP as a living document does not preclude its use in other contexts, such as for a top-level overview in a grant. In particular, this form of the DMP is effective for onboarding new researchers to a project or any time project personnel change.
Conclusion
Data management is the sum of a number of small practices that add up to being able to find and use data when you need it. Data management is most effective when these practices are habitual — consistent routines performed without extra effort. This does not mean a researcher has to totally upend her workflows to see benefits. Data management can be adopted when a significant change occurs such as a new collaboration or funding or when you get a new piece of equipment. These situations provide an excellent opportunity to take 5 minutes to determine a naming convention, for example, that can save significant time later when looking for a specific file. Alternatively, an established researcher or team could try out one new Practice per month and have better data management within a calendar year.
An incremental approach to changing data management behaviors is generally the most practical. Any changes should be discussed as a group and prioritized for implementation. Don’t try to implement all 10 Practices above all in one week, or even one month. Choose one change to adopt at a time and give people enough time to make it a habit. Once the new habit is formed, choose another to implement. In some cases, it may make more sense to significantly modify a particular workflow through multiple changes. If this approach is necessary, don’t modify all the workflows for a project at the same time. Change is stressful and new habits take time to develop. Finally, remember that when your collaborators are expressing confusion and asking “how” questions, it is usually an indication that they are open to the changes and moving toward acceptance and adoption.
Research data management may have unexpected challenges but, by addressing common issues, research teams have the opportunity to prevent data loss or misinterpretation and reduce decision fatigue. Additional planning may be needed to improve mentoring and training, performing open science, engaging with disciplinary data standards, and sharing data, yet the techniques presented here are intended to provide early success for improved processes and data capture. Evaluating current practices as reflected in data management plans is one approach to identifying where to start; a number of resources have been developed that may be useful (
Acknowledgements
The authors would like to thank Kara Woo who read and provided thoughtful comments on the manuscript. The authors acknowledge the Research Open Access Publishing (ROAAP) Fund of the University of Illinois at Chicago for financial support towards the open access publishing fee for this article.
Author contributions
Kristin Briney: Conceptualization, Project Administration, Writing – Original Draft Preparation, Writing – Review & Editing.
Heather Coates: Visualization, Writing – Original Draft Preparation, Writing – Review & Editing.
Abigail Goben: Writing – Original Draft Preparation, Writing – Review & Editing.
Conflicts of interest
The authors report no conflicts of interest.
References
-
Hard drive data and stats. https://www.backblaze.com/b2/hard-drive-test-data.html. Accessed on: 2020-2-10.
-
1,500 scientists lift the lid on reproducibility.Nature533(7604):452‑454. https://doi.org/10.1038/533452a
-
Yankaskas settles appeal, agrees to retire from UNC. https://www.dailytarheel.com/article/2011/04/yankaskas_settles_appeal_agrees_to_retire_from_unc. Accessed on: 2020-2-10.
-
Support your data: A research data management guide for researchers.Research Ideas and Outcomes4https://doi.org/10.3897/rio.4.e26439
-
How to make a data dictionary. http://help.osf.io/m/bestpractices/l/618767-how-to-make-a-data-dictionary. Accessed on: 2020-2-10.
-
Data management for researchers: organize, maintain and share your data for research success.Pelagic Publishing,Exeter, UK. [ISBN9781784270124]
-
The problem with dates: Applying ISO 8601 to research data management.Journal of eScience Librarianship7(2). https://doi.org/10.7191/jeslib.2018.1147
-
The citation advantage of linking publications to research data.arXivURL: https://arxiv.org/abs/1907.02565
-
Standards. https://www.cdisc.org/. Accessed on: 2020-6-09.
-
Guide to writing "readme" style metadata. https://data.research.cornell.edu/content/readme. Accessed on: 2020-2-10.
-
Managing and sharing research data: A guide to good practice.2.SAGE[ISBN978-1526460257]
-
Bringing citations and usage metrics together to make data count.Data Science Journal18https://doi.org/10.5334/dsj-2019-009
-
Data life cycle. https://www.dataone.org/data-life-cycle. Accessed on: 2020-2-10.
-
Overview of funders' data policies. http://www.dcc.ac.uk/resources/policy-and-legal/overview-funders-data-policies. Accessed on: 2020-3-06.
-
The ubiquitous digital file: A review of file management research.Journal of the Association for Information Science and Technology71(1):E1‑E32. https://doi.org/10.1002/asi.24222
-
Terms of Service, Curation. https://www.datadryad.org/pages/policies#curation. Accessed on: 2020-3-06.
-
Describe your dataset in a README file. https://datadryad.org/stash/best_practices#describe. Accessed on: 2020-2-10.
-
Grant reviewer's guide for data management plans. https://osf.io/SNYFB/. Accessed on: 2020-3-06.
-
Data packages. https://frictionlessdata.io/data-packages/. Accessed on: 2020-3-06.
-
Preempting discord: Prenuptial agreements for scientists. https://ori.hhs.gov/preempting-discord-prenuptial-agreements-scientists. Accessed on: 2020-2-10.
-
Electronic lab notebooks. https://datamanagement.hms.harvard.edu/electronic-lab-notebooks. Accessed on: 2020-2-10.
-
Guide to Codebooks. https://www.icpsr.umich.edu/files/deposit/Guide-to-Codebooks_v1.pdf. Accessed on: 2020-2-10.
-
ICPSR: A case study in repository management. https://www.icpsr.umich.edu/icpsrweb/content/datamanagement/lifecycle/index.html. Accessed on: 2020-3-06.
-
Guidelines for effective data management plans. https://www.icpsr.umich.edu/icpsrweb/content/datamanagement/dmp/. Accessed on: 2020-3-06.
-
Writing the laboratory notebook.American Chemical Society[ISBN978-0841209336]
-
Security Hacks. https://web.archive.org/web/20110128163359/https://www.insidehighered.com/news/2011/01/27/unc_case_highlights_debate_about_data_security_and_accountability_for_hacks. Accessed on: 2020-2-10.
-
The role of data reuse in the apprenticeship process.Proceedings of the American Society for Information Science and Technology50(1):1‑10. https://doi.org/10.1002/meet.14505001051
-
How to pick an electronic laboratory notebook.Nature560(7717):269‑270. https://doi.org/10.1038/d41586-018-05895-3
-
Recommended formats statement. https://www.loc.gov/preservation/resources/rfs/. Accessed on: 2020-3-06.
-
Team mental models and team performance: a field study of the effects of team mental model similarity and accuracy.Journal of Organizational Behavior27(4):403‑418. https://doi.org/10.1002/job.387
-
Keeping data readable in the long run. https://smallpondscience.com/2017/05/15/keeping-data-readable-in-the-long-run/. Accessed on: 2020-3-06.
-
Ten simple rules for creating a good data management plan.PLOS Computational Biology11(10):e1004525. https://doi.org/10.1371/journal.pcbi.1004525
-
Questions for scientific collaborators. https://ombudsman.nih.gov/sites/default/files/Sample%20Partnering%20Agreement%20Template.pdf. Accessed on: 2020-2-10.
-
Promoting an open research culture.Science348(6242):1422‑1425. https://doi.org/10.1126/science.aab2374
-
Managing data for integrity: Policies and procedures for ensuring the accuracy and quality of the data in the laboratory.Science and Engineering Ethics12(1):23‑39. https://doi.org/10.1007/s11948-006-0004-0
-
NASA erased first moonwalk tapes, but restores copies. https://www.space.com/6994-nasa-erased-moonwalk-tapes-restores-copies.html. Accessed on: 2020-2-10.
-
Failure trends in a large disk drive population. In:5th USENIX Conference on File and Storage Technologies. URL: https://www.usenix.org/legacy/events/fast07/tech/full_papers/pinheiro/pinheiro_old.pdf
-
Data reuse and the open data citation advantage.PeerJ1https://doi.org/10.7717/peerj.175
-
Recommended repositories. https://journals.plos.org/plosone/s/recommended-repositories. Accessed on: 2020-3-06.
-
ReproZip. https://www.reprozip.org/. Accessed on: 2020-3-06.
-
Browse data sharing requirements by Federal Agency. http://datasharing.sparcopen.org/data. Accessed on: 2020-3-06.
-
When Amazon web services goes down, so does a lot of the web. https://nymag.com/intelligencer/2018/03/when-amazon-web-services-goes-down-so-does-a-lot-of-the-web.html. Accessed on: 2020-3-06.
-
DMPTool. https://dmptool.org/. Accessed on: 2020-3-06.
-
How to start–and keep–a laboratory notebook: Policy and practical guidelines. http://www.iphandbook.org/handbook/ch08/p02/. Accessed on: 2020-2-10.
-
Research data lifecycle. https://www.ukdataservice.ac.uk/manage-data/lifecycle. Accessed on: 2020-2-10.
-
NetCDF Conventions. https://www.unidata.ucar.edu/software/netcdf/conventions.html. Accessed on: 2020-6-09.
-
DRUM services. https://conservancy.umn.edu/pages/drum/services/. Accessed on: 2020-3-06.
-
Data dictionaries. https://www.usgs.gov/products/data-and-tools/data-management/data-dictionaries. Accessed on: 2020-2-10.
-
Team learning: building shared mental models.Instructional Science39(3):283‑301. https://doi.org/10.1007/s11251-010-9128-3
-
Conceptual-level workflow modeling of scientific experiments using NMR as a case study.BMC Bioinformatics8(1). https://doi.org/10.1186/1471-2105-8-31
-
The availability of research data declines rapidly with article age.Current Biology24(1):94‑97. https://doi.org/10.1016/j.cub.2013.11.014
-
The DART Project: using data management plans as a research tool. https://doi.org/10.17605/OSF.IO/KH2Y6. Accessed on: 2020-3-06.
-
Ten simple rules for responsible big data research.PLOS Computational Biology13(3):e1005399. https://doi.org/10.1371/journal.pcbi.1005399