|
Research Ideas and Outcomes :
Grant Proposal
|
|
Corresponding author: Sonia Vanderhoeven (s.vanderhoeven@biodiversity.be), Quentin John Groom (quentin.groom@plantentuinmeise.be)
Received: 25 Apr 2017 | Published: 02 May 2017
© 2017 Sonia Vanderhoeven, Tim Adriaens, Peter Desmet, Diederik Strubbe, Thierry Backeljau, Yvan Barbier, Dimitri Brosens, Julien Cigar, Maxime Coupremanne, Rozemien De Troch, Hilde Eggermont, André Heughebaert, Kris Hostens, Pierre Huybrechts, Anne-Laure Jacquemart, Luc Lens, Arnaud Monty, Jean-Yves Paquet, Céline Prévot, Tim Robertson, Piet Termonia, Ruben Van De Kerchove, Gert Van Hoey, Bert Van Schaeybroeck, Diemer Vercayie, Thomas Verleye, Sarah Welby, Quentin Groom
This is an open access article distributed under the terms of the Creative Commons Attribution License (CC BY 4.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation:
Vanderhoeven S, Adriaens T, Desmet P, Strubbe D, Backeljau T, Barbier Y, Brosens D, Cigar J, Coupremanne M, De Troch R, Eggermont H, Heughebaert A, Hostens K, Huybrechts P, Jacquemart A, Lens L, Monty A, Paquet J, Prévot C, Robertson T, Termonia P, Van De Kerchove R, Van Hoey G, Van Schaeybroeck B, Vercayie D, Verleye T, Welby S, Groom Q (2017) Tracking Invasive Alien Species (TrIAS): Building a data-driven framework to inform policy. Research Ideas and Outcomes 3: e13414. https://doi.org/10.3897/rio.3.e13414
|
|
Abstract
Imagine a future where dynamically, from year to year, we can track the progression of alien species (AS), identify emerging problem species, assess their current and future risk and timely inform policy in a seamless data-driven workflow. One that is built on open science and open data infrastructures. By using international biodiversity standards and facilities, we would ensure interoperability, repeatability and sustainability. This would make the process adaptable to future requirements in an evolving AS policy landscape both locally and internationally. In recent years, Belgium has developed decision support tools to inform invasive alien species (IAS) policy, including information systems, early warning initiatives and risk assessment protocols. However, the current workflows from biodiversity observations to IAS science and policy are slow, not easily repeatable, and their scope is often taxonomically, spatially and temporally limited. This is mainly caused by the diversity of actors involved and the closed, fragmented nature of the sources of these biodiversity data, which leads to considerable knowledge gaps for IAS research and policy. We will leverage expertise and knowledge from nine former and current BELSPO projects and initiatives: Alien Alert, Invaxen, Diars, INPLANBEL, Alien Impact, Ensis, CORDEX.be, Speedy and the Belgian Biodiversity Platform. The project will be built on two components: 1) The establishment of a data mobilization framework for AS data from diverse data sources and 2) the development of data-driven procedures for risk evaluation based on risk modelling, risk mapping and risk assessment. We will use facilities from the Global Biodiversity Information Facility (GBIF), standards from the Biodiversity Information Standards organization (TDWG) and expertise from Lifewatch to create and facilitate a systematic workflow. Alien species data will be gathered from a large set of regional, national and international initiatives, including citizen science with a wide taxonomic scope from marine, terrestrial and freshwater environments. Observation data will be funnelled in repeatable ways to GBIF. In parallel, a Belgian checklist of AS will be established, benefiting from various taxonomic and project-based checklists foreseen for GBIF publication. The combination of the observation data and the checklist will feed indicators for the identification of emerging species; their level of invasion in Belgium; changes in their invasion status and the identification of areas and species of concern that could be impacted upon by bioinvasions. Data-driven risk evaluation of identified emerging species will be supported by niche and climate modelling and consequent risk mapping using critical climatic variables for the current and projected future climate periods at high resolution. The resulting risk maps will complement risk assessments performed with the recently developed Harmonia+ protocol to assess risks posed by emergent species to biodiversity and human, plant, and animal health. The use of open data will ensure that interested stakeholders in Belgium and abroad can make use of the information we generate. The open science ensures everyone is free to adopt and adapt the workflow for different scenarios and regions. The checklist will be used at national level, but will also serve as the Belgian reference for international databases (IUCN - GRIIS, EASIN) and impact assessments (IPBES, SEBI). The workflow will be showcased through GEO BON, the Invasivesnet network and the COST Actions Alien Challenge and ParrotNet. The observations and outcomes of risk evaluations will be used to provide science-based support for the implementation of IAS policies at the regional, federal and EU levels. The publication of Belgian data and checklists on IAS is particularly timely in light of the currently ongoing EU IAS Regulation and its implementation in Belgium. By proving that automated workflows can provide rapid and repeatable production of information, we will open up this technology for other conservation assessments.
Keywords
Information Technology; Risk Evaluation; Evidence-Based Policy; Open Data Publication; Climate Change
State of the Art and Objectives
Biogeography, dispersal biology and pest control were formerly a static study of defined distributions, local dispersal, and well-defined sets of organisms. However, globalization has forced us to think from a dynamic, long-term perspective for a whole suite of organisms, particularly with regard to AS. Human-mediated introductions of AS have become a defining feature of global environmental change (
- The establishment of a data mobilization framework for AS data from diverse data sources;
- The development of data-driven procedures for risk evaluation based on risk modelling, risk mapping and risk assessment.
Methods
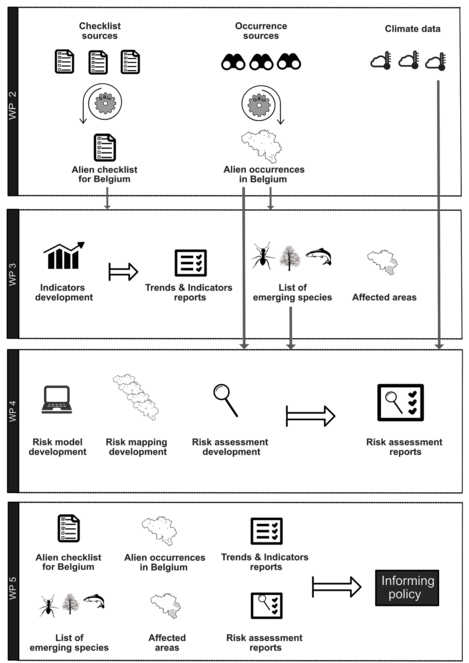
Fig.
A visual description of the TrIAS workflow through work packages. Work package 1 generates the input data; Work package 2 creates indicators and summaries of the data; Work package 3 uses the data and generates models and predications of future distributions; Work package 4 involves experts using the information from the other work packages, together with their own experience to create impact assessments.
Alien species checklist : Species’ alien status, and a priori knowledge of the presence of an alien species in a country, is central to target monitoring and control strategies (
Alien species occurrence data : Spatially and temporally explicit species occurrence records are the basic unit to track and assess range expansion of species (
Climate data and scenario : Several databases offer free access to a wide range of global bioclimatic variables, such as WorldClim (
Trends and indicators
Aichi Target 9 of the 2011–2020 Strategic Plan for Biodiversity states that “By 2020, IAS should be identified and prioritized, priority species controlled or eradicated and measures should be in place to manage invasion pathways” (
Data-driven procedures for risk evaluation
Proper decisions for IAS management depend on accurate spatial and temporal characterization of the evaluation of entry, exposure and consequence (
Risk modelling and mapping : Spatially explicit predictions of invasion risk derived from species distribution models [SDMs, also known as ecological niche models (ENMs)], calibrated with native species distributions, are increasingly incorporated into alien species risk assessments (
Risk assessment : Risk assessment protocols are tools to condense species information into their perceived risks according to a common framework. The Belgian Harmonia+ risk assessment protocol was designed during the BELSPO project Alien Alert (
Informing policy
A critical, yet often overlooked, area of risk analysis is the dissemination of information on the risks of AS introductions through risk communication. Ensuring repeatability, reliability, and transparency in both the data mobilization framework and the risk evaluation process will effectively increase understanding of the risks encountered and therefore facilitate the decision making (
Data
TrIAS will make use of existing species occurrence data (mostly observations, but also specimens) from regional administrations (INBO, ILVO & SPW-DEMNA), NGOs (Natuurpunt & Natagora), collections (BGM, RBINS), former AS related projects (Invaxen, Diars, Inplanbel, Ensis, MEMO), and AS data available on GBIF. Dedicated monitoring for AS is currently lacking in Belgium. Yet, several monitoring schemes, such as for Natura 2000, the Water Framework Directive and the Marine Strategy Framework Directive yield data on AS. Also, there are many opportunistic observations by citizen scientists. These data have never been brought together, but together they would represent the most complete and up to date source of AS occurrence data for Belgium. Where not the case already, these data will be standardized and published as open data through GBIF, so these can be used for data-driven procedures in TrIAS as well as by anyone else. In order to specifically select AS occurrence data, a verified and standardized list of all AS in Belgium is required. As such a unified checklist does currently not exist, TrIAS will make use of several authoritative checklists with a more specialized scope that collectively comprise the most complete data source for AS in Belgium. These include the Manual of Alien Plants (BGM,
Workplan and detailed description of tasks
Work Package 1 - Management
The coordinator will be responsible for the overall management of the project and the generation of the deliverables. The lead scientist in each institution will be responsible for their day-to-day task management. A formal meeting of the whole consortium will be held every six months, whereas ad hoc meetings of the task groups will be called by the task leaders or coordinator as it is deemed necessary. The whole team will work with the follow-up committee. Three meetings of the follow-up committee with the complete consortium are envisaged, one in month 1 of the project, one midway through the project and one during the final three months. The follow up committee meetings will be held at the same time as meetings of the whole consortium to reduce costs and optimise attendance. These meetings will serve to direct and consolidate the project, address problems, and get feedback from the follow-up committee. In addition, teleconference meetings and email will be used extensively to ensure coordinated working across the partners. The management will ensure progress of deliverables and milestones will be closely monitored to ensure that deadlines are met.
Task 1.1: Data Management Plan
The data management plan will describe how the information generated by the project will be handled both during and after it is generated, and will be designed to ensure that data are of good quality, standardized for better reusability, assigned appropriate metadata, and are adequately preserved (
Task 1.2: Follow-up Committee Reports
Before the second and third follow-up committee meetings a status report will be written detailing the progress of the project. This will document the use of resources and the progress towards goals. This report will also be used to report on issues related to gender and ethic within the project. Each partner will contribute to this report. They will detail their use of the budget and their individual progress towards aims. If problems are encountered during the project they will be detailed in the report together with the solutions. The report will be used to communicate progress to the follow up committee and to Belspo, but it also a useful milestone for the project to reflect on its progress.
Work Package 2: Data Mobilization Framework
The goal of the data mobilization framework is to create data products that will be used in further work packages. This includes datasets relating to the presence of AS in Belgium (checklist and occurrence data) for which major sources in Belgium will be published to GBIF, and environmental predictors needed for spatially explicit risk modelling and mapping (global and CORDEX-derived climate (change) data, habitat and land-use layers). The framework also comprises the tools and documentation to compile from the published checklists a unified AS checklist for Belgium, use this to query and download occurrence data from GBIF, and process these to feed further analysis (WP3 & WP4).
TASK 2.1 - Alien species checklist
TrIAS will work with the authors of 7 authoritative checklists of AS in Belgium (see "Data") to publish these as standardized, open datasets. Some of these checklists are one-off publications whereas others are dynamic, being continually updated. For each source, we will create a mapping scheme to standardize the original data model to the Darwin Core (DwC) standard, in large part following the GBIF Global Names Architecture (GNA) profile (
TASK 2.2 - Occurrences
As with the AS checklist, TrIAS will work with the owners of important sources of AS occurrence records for Belgium (see "Data") to publish these as standardized, open datasets. For each source, we will create a mapping schema to standardize the original data model to the Darwin Core (DwC) standard. The published occurrence data will at least include the species name, date, location and source information for each record. As with the checklists, each dataset will be documented with metadata and will be packaged as a Darwin Core Archive and registered with a DOI. We will use the GBIF Integrated Publishing Toolkit (IPT) (
TASK 2.3 - Climatic Data and Future Scenarios
The aim of this task is to develop climate prediction products targeted for enabling risk modelling (WP4) and consequent risk assessment (WP3.3) for Belgium under current and future climate conditions. While climate data needed for the ‘global’ species distribution models can be easily derived from online repositories, fine-grained Belgian-level models require tailored high-resolution data and GIS layers for current and future climates. In TriAS, such datasets will be prepared, based on and adapted from the high-resolution (4km, 16km2) results of the BELSPO-funded CORDEX.be project. These Belgian-level climate predictions will be obtained using multiple dynamically-downscaled simulations, and will explicitly include climate uncertainty estimates. Such high resolution data are required for different reasons, including a realistic representation of convective rain (
WP3 - Trends and Indicators
Tracking the spread of AS and evaluating the effectiveness of policies and management interventions is generally achieved by repeated measurement of occurrence records of AS. It implies the assessment of their geographic distribution (
TASK 3.1 - Review of existing IAS indicators
As a first step, TriAS will review existing IAS indicators in Belgium (e.g.
TASK 3.2 - Development of indicators for biological invasions in Belgium
Once the relevant indicators for invasion have been defined, TrIAS will develop threshold values for each, allowing us to detect important changes in the trends of the selected indicators. Although IAS indicators at the international level (e.g. trends in the number of IAS and the number of high impact AS) are indispensable, they are not refined enough to provide the necessary information for IAS policy and invasion management at national or regional level (e.g. early warning, risk evaluation and rapid response mechanisms). Also, at national and regional levels, information is needed focussing on certain taxa (e.g. the species of EU concern) or certain areas such as protected areas (e.g. NATURA 2000 areas, nature reserves) and entry points. The same is true for the essential variables for invasion (e.g. alien status) which primarily serve the detection of international/global trends. TrIAS will therefore also consider supplementary information such as abundance of a species, characteristics of the invaded area, pathways of introduction, and spread. We will then develop an open source tool to automatically follow trends in the defined indicators for AS in Belgium, based on public occurrence records registered with GBIF. The tool will allow for processing the occurrence data gathered in WP2, correct these for several biases and combine these with other layers of information pertinent to the indicators we want to follow, such as geospatial layers with habitat or protected area information. We will contract GBIF to assist in the development of this tool, which will potentially be built on the species-population tool developed by them (https://demo.gbif.org/tools/species-population).
TASK 3.3 - Trends and indicators report - identification of emerging alien species and affected areas
The tools developed in the previous section will be processed with occurrence data gathered in WP2. This will occur twice during the project and will report for which indicators, species and locations certain thresholds have been crossed. The indicators and threshold values defined within TrIAS will act as warnings of important changes in the status of biological invasions in Belgium, such as a change in the number of AS in Belgium, a rate of change in the distribution and/or spread of AS in Belgium, a rate of change in the distribution and/or spread of an AS in protected/priority areas of conservation concern in Belgium (
Work Package 4 - Risk evaluation
TASK 4.1 - Risk model development
Data gathering and organization
Species occurrence data: a first step in risk model development is the collection and organization of required species occurrence data. The data mobilization framework (WP2) will allow us to identify the species for which invasion risk predictions need to be generated. For the global model (i.e. the model aimed at characterizing the full range of climatic conditions under which a species can persist), global species occurrence data will be gathered from various online repositories (including those published by TrIAS). Both generic (e.g. GBIF, OBIS) and taxa-specific repositories (e.g. VertNet, AntNet,..) will be queried. Species occurrence data gathering (and dynamic updating) will be facilitated through the use of designated R packages that directly communicate with these databases (e.g. the ‘rgbif’ (
Environmental predictor variables
The global model needs spatial climate data that are available worldwide (i.e. data need to cover all areas where species occurrences are drawn from), and these data will be obtained through recognized online repositories such as WorldClim (
TASK 4.2 - Risk mapping development
Model building: spatially explicit predictions of invasion risk will be generated using an ensemble-modelling framework such as biomod2 (
Model evaluation: Predictive accuracy of models will be evaluated in the following ways. (1) Global model projections derived from native-range data only will be evaluated against invasive range occurrence data. Such invasive range occurrence data represent a truly independent dataset and allow for the strongest evaluation of model performance possible (
Model post-processing: The above-mentioned ensemble models will results in a large number of model predictions, allowing to derive, for each location (i.e. grid pixel), a probability distribution of invasion potential rather than a single crude value. This allows for extraction of average predictions, as well as confidence intervals given varying data inputs and modelling strategies. The central tendencies of model predictions for each invasive species will be obtained through (1) only considering those individual models who meet predetermined model evaluation criteria (sensu
TASK 4.3 - Risk assessment protocol development
The Harmonia+ framework was developed during the Belspo-funded Alien Alert project (
TASK 4.4 - Risk assessments
Risk assessments will be performed for emerging species identified from the application of indicators and trends (task 3.3) using the updated version of Harmonia+ (task 4.3) and the integrated risk maps as baseline information (task 4.2). Expert elicitation will involve experts from the TrIAS consortium as well as experts selected from the expert registry of the Belgian Forum on Invasive Species (http://ias.biodiversity.be/registry/index) and marine experts from the 'Alien species in the Belgian part of the North Sea and adjacent estuaries' (www.vliz.be/en/imis?module=project&proid=2170). Facilitation of the process will be done by the BBPF. Particular attention will be paid to transparency and repeatability of assessments and quality control of risk assessments. A consensus building process will be applied to capture opinions of minimum four different experts, thereby maximizing the evidential basis (
WP5 - Informing policy and stakeholders
Disseminating the information on the risk of AS is a critical part of the risk analysis process. Effective communication requires the definition of target audience, objectives, clear messages and tools to be used, and evaluation of the outcomes (
- Project website: We will use the Open Science Framework (OSF, https://osf.io/) to host a project website for TrIAS. OSF allows the documentation and communication on the project in a wiki, facilitates collaboration among the partners and external collaborators, integrates with tools such as GitHub, and provides a means to cite an ongoing open science project.
- Access to TrIAS products: Many of the outputs of the project, particularly maps and data layers, have many potential scientific applications in and beyond IAS research. These will be made available and communicated to scientists. Open source code will be deposited publicly on GitHub (https://github.com), an online code collaboration platform.
- Academic publication: For datasets/checklists published through TrIAS, data papers to peer reviewed, open access journals will be considered. Also, significant results, protocols and insights that emerge from the research will also be published in open access journals.
- Internal promotion within the institution: The project will also be promoted internally within our institutions to ensure long-term sustainability of the project. We will support and encourage the submission of additional grant proposals by our scientists who want to use this workflow to generate data for their research projects.
TASK 5.1 - Communication plan
This task will allow TrIAS to plan and coordinate communication actions. The goal of the plan is to improve the impact and visibility of TrIAS activities, by putting forward a coherent image and by identifying and focalizing on its core target and audience. Following the approach of the Belgian Biodiversity Platform, TrIAS communication will be neutral, scientifically sound, comprehensive and based on factual information. It must provide policy-relevant - but not policy-prescriptive - information; and it should ascertain equitable, unbiased representation of, and networking with and amongst, scientists and policy makers. The communication plan will include an overview of communication actions across the work packages and identify the top level messages of relevance to different target audiences.
TASK 5.2 - Risk communication
Beyond the classical risk scores allocated to species in the risk assessments, TrIAS will review and further develop innovative graphic tools to facilitate the integration of risk and related uncertainty. As such, the scientific information on Harmonia will be ready for use for policy makers and managers. A particular attention will be paid to explicit communication on the interpretation and use of risk maps and the consideration of the climate change perspective by providing detailed documentation on models and assessments development (
TASK 5.3 - Integration of TrIAS output in the national IAS information system Harmonia
The AS checklist (task 2.1), AS distribution maps (task 2.2) and the outcome of risk evaluations (task 3.3), including risk maps and graphic outputs will be integrated in the national information system Harmonia (http://ias.biodiversity.be). As a result, the main portal will be updated to allow the inclusion of these new types of outputs.
TASK 5.4 - Targeted communication actions
A particular effort will be done to 1) explain the importance of openly published data to effectively tackle IAS at several levels in the management cycle; 2) Broadcast opportunities for involvement in data publication amongst target groups; 3) Promote findings of the project widely (such as the developed automated procedures and risk evaluation tools), using appropriate mechanisms and in a format tailored to target audiences; 4) Promote project findings strategically. Two milestone communication events are planned throughout the project, geared towards different stakeholder groups and with different objectives. These include:
- A session at an international conference to showcase the results and methodology of TrIAS and gain feedback from an international audience of scientists and experts (end of year 2);
- A symposium aimed at informing the community of practice on IAS in Belgium on the outcomes of the project (year 3).
Targeted communication actions will aim at increasing the participation of the public in IAS recording. A particularly important group to reach out to are naturalist observers such as the user community of ‘waarnemingen.be’ (Natuurpunt) and ’observations.be’ (Natagora), the marine citizen science initiative "SeaWatch-B" (VLIZ, http://www.seawatch-b.be/), or official portals such as the regional institutional portal for biodiversity observations (http://observatoire.biodiversite.wallonie.be/encodage). We would like to encourage recording activity on AS.
Expected impacts of the research and compliance of the research with the expected impacts
Expected scientific impacts / research community
IAS are an international problem and as such require cooperative international action. By basing TrIAS on the tenets of open science and in global repositories we facilitate an international approach and will lead the way in providing scientific basis for decision-making in conservation management. Results of our analyses will be published in international open access journals. Also, in the medium term, additional observations from Belgium will help other countries model and assess the risks of these invasive species under different future scenarios. In the long term, the consolidation, standardization and openness of data are likely to lead to novel scientific applications of those data and workflows that go beyond what TrIAS is proposing.
Expected impacts on policy support / policy makers
Much has been written about the importance of evidence-based decision making, however policy makers still find themselves with inadequate, contradictory information. This information can be out of date, not communicated clearly and advice may lack clear expressions of certainty. This is as true for information on IAS as it is for many other policy areas. There are several causes of this, but they include the fragmented ownership of data resources; a lack of information technology infrastructure and the lack standardization. TrIAS will address these issues by creating a rapid, joined-up and sustainable way to inform policy on IAS in Belgium. The time it takes from the observation of an organism in the field, to that observation being used policy will be reduced from years to months.
Expected societal impacts
TrIAS will address goals 9 and 15 of the sustainable development agenda of the UN. These are specifically to “build a resilient infrastructure and foster innovation”, but also to protect biodiversity and sustainable land-use. Controlling and potentially eradicating AS is expensive, however, these costs can be reduced significantly through rapid targeted actions. The dramatic reduction in the time from the collection of observations to the creation of actionable evidence should have a positive impact on the outcomes of management actions, by speeding reaction times and reducing wasted effort. Such outcomes will also have positive benefits for the reduction of animal suffering during management and for the reduction of coincidental damage to ecosystems due to management actions on IAS. An essential element in the whole information flow are the volunteers and professionals who collect and collate observations and checklists in Belgium. This project will valorize their effort to improve knowledge and conserve Belgium’s natural heritage. We can expect that TrIAS will encourage their work by showing them how their observations help conserve biodiversity. Furthermore, TrIAS will bring the work of these organizations to the attention of policy makers, making them more aware of the vital role these organizations have in monitoring biodiversity. By doing so, we expect additional societal benefits such as improved science literacy among participants and a greater understanding and awareness of the invasive species issue.
Funding program
This was a proposal to the Belgian Science Policy Office call for Belgian Research Action through Interdisciplinary Networks (BRAIN).
References
- Ensemble forecasting of species distributions.Trends in Ecology & Evolution22(1):42‑47. https://doi.org/10.1016/j.tree.2006.09.010
- Five (or so) challenges for species distribution modelling.Journal of Biogeography33(10):1677‑1688. https://doi.org/10.1111/j.1365-2699.2006.01584.x
- Uses and misuses of bioclimatic envelope modeling.Ecology93(7):1527‑1539. https://doi.org/10.1890/11-1930.1
- Improving species distribution models for climate change studies: variable selection and scale.Journal of Biogeography38(1):1‑8. https://doi.org/10.1111/j.1365-2699.2010.02416.x
- The fate of European breeding birds under climate, land-use and dispersal scenarios.Global Change Biology18:881‑890. https://doi.org/10.1111/j.1365-2486.2011.02552.x
- Selecting pseudo-absences for species distribution models: how, where and how many?Methods in Ecology and Evolution3(2):327‑338. https://doi.org/10.1111/j.2041-210x.2011.00172.x
- How can knowledge of the climate niche inform the weed risk assessment process? A case study of Chrysanthemoides monilifera in Australia.Diversity and Distributions20(6):613‑625. https://doi.org/10.1111/ddi.12190
- Spatial bias in the GBIF database and its effect on modeling species' geographic distributions.Ecological Informatics19:10‑15. https://doi.org/10.1016/j.ecoinf.2013.11.002
- Will climate change promote future invasions?Global Change Biology19:3740‑3748. https://doi.org/10.1111/gcb.12344
- Streamlining European biodiversity indicators 2020: Building a future on lessons learnt from the SEBI 2010 process.European Environment Agency,Copenhagen,45pp. [ISBN978-92-9213-326-9] https://doi.org/10.2800/55751
- Distorted Views of Biodiversity: Spatial and Temporal Bias in Species Occurrence Data.PLoS Biology8(6):e1000385. https://doi.org/10.1371/journal.pbio.1000385
- Alien macroinvertebrates in Flanders (Belgium).Aquatic Invasions11(2):131‑144. https://doi.org/10.3391/ai.2016.11.2.03
- Predicting current and future biological invasions: both native and invaded ranges matter.Biology Letters4(5):585‑589. https://doi.org/10.1098/rsbl.2008.0254
- Evidence of climatic niche shift during biological invasion.Ecology Letters10(8):701‑709. https://doi.org/10.1111/j.1461-0248.2007.01060.x
- How to communicate on pests and invasive alien plants? Conclusions of the EPPO/CoE/IUCN- ISSG/DGAV/UC/ESAC Workshop.EPPO Bulletin44(2):205‑211. https://doi.org/10.1111/epp.12110
- Uncertainty in ensemble forecasting of species distribution.Global Change Biology16(4):1145‑1157. https://doi.org/10.1111/j.1365-2486.2009.02000.x
- Strategic Plan for Biodiversity 2011-2020, including Aichi Biodiversity Targets. https://www.cbd.int/sp/targets/. Accessed on: 2017-3-11.
- Interface to the Global 'Biodiversity' Information Facility 'API'.0.9.7.CRAN. Release date:2017-1-21.
- spocc: Interface to Species Occurrence Data Sources.0.6.0.CRAN. Release date:2016-12-07. URL: https://CRAN.R-project.org/package=spocc
- Biodiversity Indicators. State of Nature in Flanders. Mededeling van het Instituut voor Natuur- en Bosonderzoek. ,Brussels,56pp. URL: https://data.inbo.be/purews/files/11365385/BiodiversityIndicators_2015.pdf
- Multiscale Performance of the ALARO-0 Model for Simulating Extreme Summer Precipitation Climatology in Belgium.Journal of Climate26(22):8895‑8915. https://doi.org/10.1175/jcli-d-12-00844.1
- Harmonia+ and Pandora+ : risk screening tools for potentially invasive organisms.Belgian Biodiversity Platform, Brussels63[Inen]. URL: http://ias.biodiversity.be/harmoniaplus
- Harmonia + and Pandora +: risk screening tools for potentially invasive plants, animals and their pathogens.Biological Invasions17(6):1869‑1883. https://doi.org/10.1007/s10530-015-0843-1
- The Use of Climatic Niches in Screening Procedures for Introduced Species to Evaluate Risk of Spread: A Case with the American Eastern Grey Squirrel.PLoS ONE8(7):e66559. https://doi.org/10.1371/journal.pone.0066559
- Modelling distribution in European stream macroinvertebrates under future climates.Global Change Biology19:752‑762. https://doi.org/10.1111/gcb.12107
- Correcting sample selection bias in maximum entropy density estimation.Advances in neural information processing systems,17.Advances in neural information processing systems,Vancouver,2004.MIT Press,323–330pp. [ISBN9780262195348].
- Climate Extremes: Observations, Modeling, and Impacts.Science289(5487):2068‑2074. https://doi.org/10.1126/science.289.5487.2068
- The art of modelling range-shifting species.Methods in Ecology and Evolution1(4):330‑342. https://doi.org/10.1111/j.2041-210x.2010.00036.x
- New or interesting lichens and lichenicolous fungi from Belgium, Luxembourg and northern France. XI.Bulletin de la Société des naturalistes luxembourgeois109:35‑51. URL: http://snl.lu/publications/bulletin/SNL_2008_109_035_051.pdf
- Our life insurance, our natural capital: an EU biodiversity strategy to 2020.European Commission,Brussels,17pp. URL: http://eur-lex.europa.eu/legal-content/EN/TXT/?uri=CELEX:52011DC0244
- Streamlining European biodiversity indicators 2020: Building a future on lessons learnt from the SEBI 2010 process. EEA Technical report No 11/2012.European Environment Agency,Copenhagen,45pp. [InEnglish]. [ISBN978-92-9213-326-9] https://doi.org/10.2800/55751
- Assessment of alternative approaches for bioclimatic modeling with special emphasis on the Mahalanobis distance.Ecological Modelling160:115‑130. https://doi.org/10.1016/s0304-3800(02)00327-7
- The projection of species distribution models and the problem of non-analog climate.Biodiversity and Conservation18(8):2255‑2261. https://doi.org/10.1007/s10531-009-9584-8
- Mapping Species Distributions with MAXENT Using a Geographically Biased Sample of Presence Data: A Performance Assessment of Methods for Correcting Sampling Bias.PLOS ONE9(5):97122. https://doi.org/10.1371/journal.pone.0097122
- Moving beyond static species distribution models in support of conservation biogeography.Diversity and Distributions16(3):321‑330. https://doi.org/10.1111/j.1472-4642.2010.00641.x
- Targeting and Prioritisation for INS in the RINSE Project Area.University ofCambridge,Cambridge, UK,98pp. [InEnglish]. URL: http://www.rinse-europe.eu/assets/__files/rinse-wp1-report-en-.pdf
- Trans-national horizon scanning for invasive non-native species: a case study in western Europe.Biological Invasions18(1):17‑30. https://doi.org/10.1007/s10530-015-0986-0
- Invasive species distribution models - how violating the equilibrium assumption can create new insights.Global Ecology and Biogeography21(11):1126‑1136. https://doi.org/10.1111/j.1466-8238.2012.00768.x
- How Large Is a Species' Geographic Range?Oikos61(3):434. https://doi.org/10.2307/3545251
- GBIF GNA Profile Reference Guide for Darwin Core Archives, version 1.2, released on 15 March 2012 (contributed by Remsen D.P., Döring, M, Robertson, T.).Global Biodiversity Information Facility,Copenhagen,28pp. [InEnglish]. URL: http://links.gbif.org/gbif_gna_profile_reference_guide [ISBN87-92020-25-9]
- GBIF Backbone Taxonomy.GBIF Secretariat. Release date:2016-7-25. URL: http://doi.org/10.15468/39omei
- Validation of the ALARO-0 model within the EURO-CORDEX framework.Geoscientific Model Development9(3):1143‑1152. https://doi.org/10.5194/gmd-9-1143-2016
- The importance of open data for invasive alien species research, policy and management.Management of Biological Invasions6(2):119‑125. https://doi.org/10.3391/mbi.2015.6.2.02
- Predictive habitat distribution models in ecology.Ecological Modelling135:147‑186. https://doi.org/10.1016/s0304-3800(00)00354-9
- Conceptual clarity, scientific rigour and 'The Stories We Are': engaging with two challenges to the objectivity of invasion biology.Fifty Years of Invasion Ecology: The Legacy of Charles Elton.John Wiley & Sons,432pp. [ISBN1444335855].
- Cross-validation of species distribution models: removing spatial sorting bias and calibration with a null model.Ecology93(3):679‑688. https://doi.org/10.1890/11-0826.1
- Very high resolution interpolated climate surfaces for global land areas.International Journal of Climatology25(15):1965‑1978. https://doi.org/10.1002/joc.1276
- Tools for visualizing and integrating pest risk assessment ratings and uncertainties*.EPPO Bulletin42(1):35‑41. https://doi.org/10.1111/j.1365-2338.2012.02548.x
- Climatologies at high resolution for the earth's land surface areas.2.arxiv.org. Release date:2016-9-21. URL: https://arxiv.org/abs/1607.00217v2
- Zeneto,s A, Zervou S European Alien Species Information Network (EASIN): supporting European policies and scientific research.Management of Biological Invasions.6,2.147-157pp. https://doi.org/10.3391/mbi.2015.6.2.05
- Regional climate modeling on European scales: a joint standard evaluation of the EURO-CORDEX RCM ensemble.Geoscientific Model Development7(4):1297‑1333. https://doi.org/10.5194/gmd-7-1297-2014
- Content analysis: An introduction to its methodology.3.Sage,441pp. [ISBN9781412983150]
- CliMond: global high-resolution historical and future scenario climate surfaces for bioclimatic modelling.Methods in Ecology and Evolution3(1):53‑64. https://doi.org/10.1111/j.2041-210x.2011.00134.x
- A vision for global monitoring of biological invasions.Biological Conservationhttps://doi.org/10.1016/j.biocon.2016.06.013
- Indicators from the global and sub-global Millennium Ecosystem Assessments: An analysis and next steps.Ecological Indicators17:77‑87. https://doi.org/10.1016/j.ecolind.2011.04.025
- The uncertain nature of absences and their importance in species distribution modelling.Ecography33(1):103‑114. https://doi.org/10.1111/j.1600-0587.2009.06039.x
- BIOLOGICAL INVASIONS: RECOMMENDATIONS FOR U.S. POLICY AND MANAGEMENT.Ecological Applications16(6):2035‑2054. https://doi.org/10.1890/1051-0761(2006)016[2035:birfup]2.0.co;2
- Risk assessment as a tool for evaluating risk management options for food safety.Towards a risk-based chain control.4.Wageningen Academic Publishers,Wageningen,408pp. [ISBN978-90-76998-97-8]. https://doi.org/10.3920/978-90-8686-583-3
- Projecting future expansion of invasive species: comparing and improving methodologies for species distribution modeling.Global Change Biology21(12):4464‑4480. https://doi.org/10.1111/gcb.13038
- Characterizing common and range expanding species.Journal of Biogeography43(2):217‑228. https://doi.org/10.1111/jbi.12642
- Uncertainty in invasive alien species listing.Ecological Applications22(3):959‑971. https://doi.org/10.1890/11-1252.1
- An Essential Biodiversity Variable approach to monitoring biological invasions: Guide for Countries.2.GEO BON Technical Series,13pp. URL: http://www.geobon.org/Downloads/reports/GEOBON/2015/MonitoringBiologicalInvasions.pdf
- Data Curation: A Study of Researcher Practices and Needs.portal: Libraries and the Academy14(2):139‑164. https://doi.org/10.1353/pla.2014.0009
- Multidimensional biases, gaps and uncertainties in global plant occurrence information.Ecology Letters19(8):992‑1006. https://doi.org/10.1111/ele.12624
- The Delphi technique in ecology and biological conservation: applications and guidelines.Methods in Ecology and Evolution6(9):1097‑1109. https://doi.org/10.1111/2041-210x.12387
- sdm: a reproducible and extensible R platform for species distribution modelling.Ecography39(4):368‑375. https://doi.org/10.1111/ecog.01881
- Constraints on interpretation of ecological niche models by limited environmental ranges on calibration areas.Ecological Modelling263:10‑18. https://doi.org/10.1016/j.ecolmodel.2013.04.011
- World Register of Introduced Marine Species (WRIMS). http://www.marinespecies.org/introduced. Accessed on: 2016-8-26.
- Predicting the impacts of climate change on the distribution of species: are bioclimate envelope models useful?Global Ecology and Biogeography12:361‑371. https://doi.org/10.1046/j.1466-822X.2003.00042.x
- Essential Biodiversity Variables.Science339(6117):277‑278. https://doi.org/10.1126/science.1229931
- Climatic Niche Shifts Are Rare Among Terrestrial Plant Invaders.Science335(6074):1344‑1348. https://doi.org/10.1126/science.1215933
- Maximum entropy modeling of species geographic distributions.Ecological Modelling190:231‑259. https://doi.org/10.1016/j.ecolmodel.2005.03.026
- Sample selection bias and presence-only distribution models: implications for background and pseudo-absence data.Ecological Applications19(1):181‑197. https://doi.org/10.1890/07-2153.1
- Precipitation in the EURO-CORDEX 0.11° and 0.44° simulations: high resolution, high benefits?Climate Dynamics46:383‑412. https://doi.org/10.1007/s00382-015-2589-y
- No silver bullets in correlative ecological niche modelling: insights from testing among many potential algorithms for niche estimation.Methods in Ecology and Evolution6(10):1126‑1136. https://doi.org/10.1111/2041-210x.12397
- Invasive alien species indicators in Europe - A review of streamlining European biodiversity (SEBI) Indicator 10.European Environment Agency,Copenhagen,44pp. [InEnglish]. [ISBN978-92-9213-342-9] https://doi.org/10.2800/64181
- Developing and testing alien species indicators for Europe.Journal for Nature Conservation29:89‑96. https://doi.org/10.1016/j.jnc.2015.12.001
- Are niche-based species distribution models transferable in space?Journal of Biogeography33(10):1689‑1703. https://doi.org/10.1111/j.1365-2699.2006.01466.x
- The GBIF Integrated Publishing Toolkit: Facilitating the Efficient Publishing of Biodiversity Data on the Internet.PLoS ONE9(8):e102623. https://doi.org/10.1371/journal.pone.0102623
- The Assessment of African Swine Fever Virus Risk to Belgium Early 2014, using the Quick and Semiquantitative Pandora Screening Protocol.Transboundary and Emerging Diseases64:237‑249. https://doi.org/10.1111/tbed.12365
- Environmental Factors Associated with Avian Distributional Boundaries.Journal of Biogeography15(3):489‑505. https://doi.org/10.2307/2845278
- GB Non-native Species Information Portal: documenting the arrival of non-native species in Britain.Biological Invasions16(12):2495‑2505. https://doi.org/10.1007/s10530-014-0687-0
- Checklist of the Bryophytes of Belgium.Journal of Botany134:97‑120. URL: http://www.jstor.org/stable/20794485
- Niche shift in four non-native estrildid finches and implications for species distribution models.Ibis157(1):75‑90. https://doi.org/10.1111/ibi.12194
- biomod2: Ensemble Platform for Species Distribution.3.3-7.CRAN. Release date:2016-3-01. URL: ftp://ftp2.de.freebsd.org/pub/misc/cran/web/packages/biomod2/biomod2.pdf
- A mid-term analysis of progress toward international biodiversity targets.Science346(6206):241‑244. https://doi.org/10.1126/science.1257484
- SpeciesGeoCoder: Fast Categorization of Species Occurrences for Analyses of Biodiversity, Biogeography, Ecology, and Evolution.Systematic Biologysyw064. https://doi.org/10.1093/sysbio/syw064
- Equilibrium or not? Modelling potential distribution of invasive species in different stages of invasion.Diversity and Distributions18(1):73‑83. https://doi.org/10.1111/j.1472-4642.2011.00854.x
- Niet-inheemse soorten van het Belgische deel van de Noordzee en aanpalende estuaria.VLIZ Species Publication,59.Vlaams Instituut vorr de Zee,OOstende,372pp. [InDutch].
- A science-based approach to tackle invasive alien species in Belgium – the role of the ISEIA protocol and the Harmonia information system as decision support tools.Management of Biological Invasions6(2):197‑208. https://doi.org/10.3391/mbi.2015.6.2.10
- Beyond protocols: improving the reliability of expert-based risk analysis underpinning invasive species policies.Biological Invasions.
- Selecting pseudo-absence data for presence-only distribution modeling: How far should you stray from what you know?Ecological Modelling220(4):589‑594. https://doi.org/10.1016/j.ecolmodel.2008.11.010
- Catalogue des Uredinales de Belgique, 1re partie, Chaconiaceae, Coleosporiaceae, Cronartiaceae, Melampsoraceae, Phragmidiaceae, Pucciniastraceae, Raveneliaceae et Uropyxidaceae.Lejeunia, Revue de Botanique183.
- CATALOGUE DES USTILAGINALES S.L. DE Belgique.Lejeunia, Revue de Botanique193URL: http://popups.ulg.ac.be/0457-4184/index.php?id=1150
- Global exchange and accumulation of non-native plants.Nature525(7567):100‑103. https://doi.org/10.1038/nature14910
- Reliability of regional climate model trends.Environmental Research Letters8(1):014055. https://doi.org/10.1088/1748-9326/8/1/014055
- The non-indigenous ctenophore Mnemiopsis leidyi in the southern North Sea: Ecological and socio-economic effects related to its trophic position and the current distribution of gelatinous zooplankton.Institute for Agricultural and Fisheries Research (ILVO),Ghent,287pp. URL: http://pure.ilvo.vlaanderen.be/portal/files/3783423/Reviewed_PhD_thesis_Lies_Vansteenbrugge.pdf [ISBN978-90-5989-837-0].
- Species distribution modelling as a macroecological tool: a case study using New World amphibians.Ecography35(6):539‑548. https://doi.org/10.1111/j.1600-0587.2011.07050.x
- Pest Risk Maps for Invasive Alien Species: A Roadmap for Improvement.BioScience60(5):349‑362. https://doi.org/10.1525/bio.2010.60.5.5
- Assessing the quality of pest risk models.Pest Risk Modelling and Mapping for Invasive Alien Species.7.CAB books,268pp. [ISBN9781780643946].
- Catalogue of neophytes in Belgium (1800-2005).Scripta Botanica Belgica.39.National Botanic Garden,Meise, Belgium,89pp. [InEnglish]. URL: http://alienplantsbelgium.be/sites/alienplantsbelgium.be/files/tabel_2.pdf [ISBN90-72619-71-4].
- The Manual of the Alien Plants of Belgium. http://alienplantsbelgium.be. Accessed on: 2017-2-22.
- The non-indigenous freshwater fishes of Flanders (Belgium): review, status and trends over the last decade.Journal of Fish Biology71:160‑172. https://doi.org/10.1111/j.1095-8649.2007.01679.x
- Registry of non-native species in the Two Seas region countries (Great Britain, France, Belgium and the Netherlands).NeoBiota23:65‑80. https://doi.org/10.3897/neobiota.23.5665