|
Research Ideas and Outcomes : Project Report
|
|
Corresponding author: Kaori L Ito (kaoriito@usc.edu)
Received: 13 Feb 2017 | Published: 14 Feb 2017
© 2017 Kaori Ito, Julia Anglin, Sook-Lei Liew
This is an open access article distributed under the terms of the Creative Commons Attribution License (CC BY 4.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Ito K, Anglin J, Liew S (2017) Semi-automated Robust Quantification of Lesions (SRQL) Toolbox. Research Ideas and Outcomes 3: e12259. https://doi.org/10.3897/rio.3.e12259
|

|
Abstract
Quantifying lesions in a robust manner is fundamental for studying the effects of neuroanatomical changes in the post-stroke brain on recovery. However, the wide variability in lesion characteristics across individuals makes manual lesion segmentation a challenging and often subjective process. This makes it difficult to combine stroke lesion data across multiple research sites, due to subjective differences in how lesions may be defined. We developed the Semi-automated Robust Quantification of Lesions (SRQL; https://github.com/npnl/SRQL; DOI: 10.5281/zenodo.267213) Toolbox that performs several analysis steps: 1) a white matter intensity correction that removes healthy white matter voxels from the lesion mask, thereby making lesions slightly more robust to subjective errors; 2) an automated report of descriptive statistics on lesions for simplified comparison between or across groups, and 3) an option to perform analyses in both native and standard space to facilitate analyses in either space, or comparisons between spaces. Here, we describe the methods implemented in the toolbox and demonstrate the outputs of the SRQL toolbox.
Keywords
stroke, neuroimaging, software, neuroanatomy, robust, MRI, analysis
Introduction
In stroke neuroimaging research, robust lesion quantification is important for performing statistical analyses relating brain changes to behavior after stroke. Specific lesion characteristics (e.g., stroke volume and location) can provide information about inter-individual differences in post-stroke neuroanatomy. However, as conventional practice for lesion segmentation involves manually tracing lesions from structural brain images, potential inconsistencies can arise due to subjective differences in the way lesions are defined. This would be particularly problematic for large, multisite collaborative efforts that involve many different individuals segmenting lesions. Thus, we aimed to develop a robust, user-friendly toolbox to improve standardization of lesion segmentation and to capture lesion characteristics for statistical analysis across individuals.
Method
The SRQL toolbox incorporates three important features to make lesion analyses more robust: (1) a semi-automated white matter intensity correction, (2) a report of descriptive statistics on lesions, and (3) an option to perform analyses in native or standard space.
White Matter Intensity Correction
A semi-automated white matter intensity correction was implemented to remove healthy white matter voxels that may have accidentally been included in manual lesion segmentations (see Fig.
For each individual subject, a white matter mask is manually created in addition to the lesion segmentation. The white matter mask is a small circular region of interest (ROI) manually drawn on three contiguous slices within the healthy white matter of the contralesional hemisphere. Both the white matter and lesion masks are then converted into binarized files. The binarized white matter mask is overlayed onto the subject's T1 structural file to obtain a mean and standard deviation of the image intensity value within the region of the white matter mask. Next, the intensity of the subject's T1 structural file is applied onto the binarized lesion mask to create a T1-weighted lesion mask. The value of the mean plus or minus one standard deviation of the white matter intensity is calculated, and applied as threshold values on the T1-weighted lesion mask. This removes voxels within the lesion mask that are within the normal intensity range of healthy white matter for that individual. Lastly, the white matter adjusted lesion mask within the subject's file is binarized as the final lesion mask for that subject.
Descriptive Lesion Statistics
The SRQL toolbox additionally outputs a single descriptive lesion database file, organized by dataset (group) and subject. For each subject, the following descriptive statistics on white matter corrected lesions are reported and stored into the lesion database file: the hemisphere of the lesion (left or right), the lesion volume (mm3), the total brain volume (mm3), and the percentage of lesion volume to total brain volume. Specifically, the hemisphere of the lesion is estimated by extracting the x-coordinate of the center of gravity of the lesion (where a positive x-coordinate indicates a right hemisphere lesion, and a negative value indicates a left hemisphere lesion). The total lesion volume, total (skull-stripped) brain volume, and percentage of lesion-to-brain volume are then calculated using FSL's fslstats (
Registration to Standard Space
Finally, we implemented an option to convert the outputs (the adjusted lesion mask file and descriptive statistics) into standard space. As registration to standard space is computationally expensive, we elected to make this an optional feature for users who wish to compare lesion data across multiple subjects. To do this, the toolbox uses FSL's FLIRT toolbox (
Results
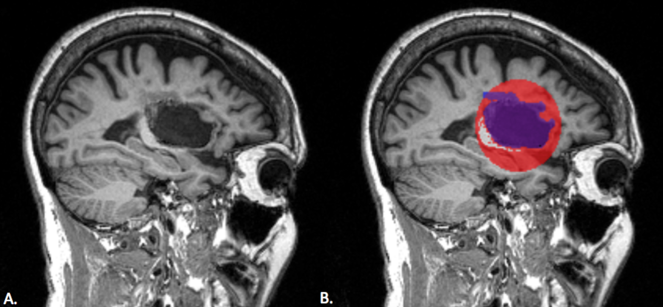
We first tested the toolbox on a single stroke subject's anatomical scan (Fig.
We tested our toolbox on a mock lesion mask. A. The stroke subject's T1 anatomical scan; B. The mock lesion mask is the red sphere; the blue mask is the lesion segmentation. The white matter was intentionally covered within the mock lesion mask, but as shown here, white matter voxels are removed by the white matter correction.
We then tested the SRQL toolbox on datasets from two different sites consisting of sixty subjects in total. Our script successfully ran on the two datasets, and we manually checked the hemispheric categorization of each lesion. We also checked the total number of voxels removed from the original lesion volume on average, which was 4.19±12.19%. Put another way, for each brain, there were approximately 4% of voxels that fell within the healthy white matter range that were subsequently removed.
Conclusion
We developed a toolbox to optimize quantification of lesions across research sites. The toolbox offers a robust analysis pipeline for performing lesion analyses across multiple datasets. Specifically, this toolbox improves the performance of statistical analysis on lesions through standardizing lesion masks with white matter adjustment, reporting descriptive lesion statistics, and normalizing adjusted lesion masks to standard space.
Future directions for the toolbox include: (1) automating the creation of each subject's white matter mask to decrease intensive labor demands and (2) constructing a report of group statistics on lesion data as well as a group probability map for visual identification of lesions across individuals.
Acknowledgements
The authors would like to thank the organizers and attendees of Brainhack LA, our research participants, and collaborators for their contributions.
Author contributions
KLI, JMA, & S-LL developed the software, performed tests, and wrote the report.
Conflicts of interest
We have no conflicts of interest to declare.
References
-
A global optimisation method for robust affine registration of brain images.Medical Image Analysis5(2):143‑156. https://doi.org/10.1016/s1361-8415(01)00036-6
-
Improved Optimization for the Robust and Accurate Linear Registration and Motion Correction of Brain Images.NeuroImage17(2):825‑841. https://doi.org/10.1006/nimg.2002.1132
-
FSL.NeuroImage62(2):782‑790. https://doi.org/10.1016/j.neuroimage.2011.09.015
-
Anatomy of Stroke Injury Predicts Gains From Therapy.Stroke42(2):421‑426. https://doi.org/10.1161/strokeaha.110.599340